Abstract
Glioblastoma (GBM) is characterized by profound immunosuppression and poor response to immunotherapy. Sex biased incidence and outcomes suggest that immune programs may be differentially regulated in males and females, but the molecular substrates of this divergence remain incompletely defined. TACSTD2 (TROP2), an epithelial lineage marker and therapeutic target in multiple solid tumors, is normally silent in the brain yet becomes aberrantly reactivated in a subset of GBM via promoter CpG hypomethylation with sex modulated prognostic effects. The immune context of TACSTD2 reactivation in GBM is unknown.
Here, we used publicly available deconvolution tools to characterize the association between TACSTD2 expression and tumor infiltrating immune cells in TCGA GBM and to place TACSTD2 within broader biological pathways through co expression and enrichment analysis. Immune infiltration was estimated using TIMER3 across multiple algorithms (including CIBERSORT and TME), and TACSTD2 correlated genes were analyzed with Enrichr (KEGG 2021 Human, MSigDB Hallmark 2020).
TIMER3 demonstrated modest but statistically significant positive correlations between TACSTD2 expression and CD4+ T cell memory estimates in GBM (CIBERSORT CD4+ memory resting: rho≈0.26, p≈2×10-3; T cell CD4+ CONSENSUS_TME: rho≈0.24, p≈4.5×10-3), with effect sizes comparable to those reported for established immune modulatory genes in deconvolution based GBM analyses. These correlations are moderate in magnitude and are interpreted as a correlative trend rather than a strong biomarker relationship. No strong negative correlation with global CD8+ T cell infiltration was observed, indicating that TACSTD2 reactivation does not simply mark a CD8 excluded or immune cold phenotype.
Pathway enrichment of TACSTD2 correlated genes (top 100, r>0.49) revealed dominant estrogen signaling, estrogen response, and apical junction signatures, alongside enrichment for ECM receptor interaction, focal adhesion and cell adhesion molecules, with minimal overlap with canonical immune checkpoint pathways. These findings support a model in which TACSTD2 marks a hormone tuned epithelial junction program that co varies with relative enrichment of CD4+ memory T cell infiltration, rather than functioning as a direct T cell intrinsic immune checkpoint. This purely computational study provides hypothesis generating, correlative evidence linking sex modulated TACSTD2 reactivation, epithelial hormone programs, and adaptive immune context in GBM, and it motivates future single cell, spatial, and functional studies to define causality and therapeutic relevance.
Keywords
Glioblastoma, TACSTD2, TROP2, Immune infiltration, CD4+ memory T cells, Estrogen signaling, Epithelial programs
Introduction
Glioblastoma (GBM) is the most aggressive primary malignant brain tumor in adults, with median survival of approximately 15 months despite maximal surgery, radiotherapy and temozolomide chemotherapy [1,2]. Profound local and systemic immunosuppression, driven by tumor-intrinsic factors and the brain microenvironment, is a hallmark of GBM and contributes to the limited efficacy of immune checkpoint blockade compared with other solid cancers [1,3]. Tumor-infiltrating lymphocytes, including CD8+ cytotoxic T cells and CD4+ memory and tissue-resident populations, have nonetheless been detected in GBM, where they can exert context-dependent antitumor or protumor functions.
Epidemiologic and clinical data indicate robust sexual dimorphism in GBM: males develop GBM more frequently and have worse outcomes than females, independent of standard prognostic factors. These observations have prompted interest in sex hormones, sex-biased immune regulation, and epigenetic control as contributors to GBM biology. Prior work has implicated sex-modulated DNA methylation at specific CpG sites as a regulator of lineage and immune programs in GBM, but mechanistic links remain sparsely defined.
TACSTD2 (tumor-associated calcium signal transducer 2; TROP2) is a transmembrane glycoprotein highly expressed in epithelia and overexpressed in multiple carcinomas, where it can drive proliferation, invasion, and therapeutic resistance. Experimental studies in epithelial malignancies, including breast cancer, have identified a TROP2–claudin tight-junction module that mediates immune exclusion and impairs CD8+ T-cell infiltration and response to PD-1 blockade, positioning TROP2 as a potential immune-modulatory barrier in that context. In contrast, TACSTD2 is nearly absent from the normal brain but becomes aberrantly reactivated in a subset of GBM through promoter CpG hypomethylation, with prior methylation analyses showing strong sex-dependent prognostic effects at the TACSTD2 locus [1,4–6].
The immune context of TACSTD2 reactivation in GBM, however, remains unknown [7,8]. It is unclear whether TACSTD2-high tumors in the brain recapitulate the immune-cold, CD8-excluded phenotype described for TROP2-high epithelial cancers, or whether lineage and tissue context fundamentally alter the immune associations of TACSTD2. Moreover, the relationship between sex-modulated TACSTD2 methylation, hormone-responsive epithelial programs and the adaptive immune milieu has not been explored in an integrative manner.
This study uses TIMER3 and complementary public resources to (i) quantify associations between TACSTD2 expression and inferred immune cell infiltration in TCGA-GBM across multiple deconvolution algorithms, and (ii) place TACSTD2 within broader immune- and hormone-relevant pathways through co-expression and enrichment analysis. The work is entirely computational and based on bulk RNA-seq deconvolution, and is therefore intended to provide modest-effect, correlative, hypothesis-generating insight, particularly regarding CD4+ memory T-cell infiltration and epithelial-hormone junction programs rather than to define a definitive prognostic or predictive biomarker. Recent work has defined immune cell signatures that stratify GBM prognosis and can guide therapy after radiotherapy [2,9].
Methods
Data sources
All analyses used publicly available datasets and tools. TCGA-GBM bulk RNA-seq and corresponding clinical data were accessed indirectly through the TIMER3 and GEPIA2 web platforms, which provide pre-processed expression matrices and deconvolution outputs. No new patient data were generated, and no additional ethical approval was required.
TACSTD2 expression (log2 TPM) values for TCGA-GBM tumors were obtained within TIMER3 (Immune_Gene module) and GEPIA2. Analyses were restricted to primary GBM samples with complete expression profiles (n≈287 in TIMER3). For pan-cancer visualization of TACSTD2 expression, tumor and matched normal TPM values across TCGA and GTEx datasets were accessed via GEPIA2, using default settings [10].
Immune infiltration analysis (TIMER3)
The TIMER3 Immune_Gene module was used to evaluate associations between TACSTD2 expression and estimated abundance of tumor-infiltrating immune cell subsets in GBM. The following settings were applied:
- Gene: TACSTD2
- Cancer type: GBM (TCGA)
- Immune infiltrates: default panel, including CD4+ and CD8+ T-cell subsets, B cells, NK cells, macrophages, dendritic cells, and myeloid-derived populations
- Algorithms: multiple deconvolution methods (CIBERSORT, CIBERSORT-ABS, xCell, quanTIseq, TME, EPIC, and others available in TIMER3)
TIMER3 reports tumor-purity-adjusted Spearman partial correlation coefficients (rho) between gene expression and inferred immune cell abundance for each algorithm. These results were exported as a heatmap of partial correlations across all immune subsets and deconvolution methods (Figure 1), with significance (p ≤0.05) indicated in the tool-generated output. TIMER3 builds on the TIMER framework for estimating tumor-infiltrating immune cells from bulk RNA-seq [10].
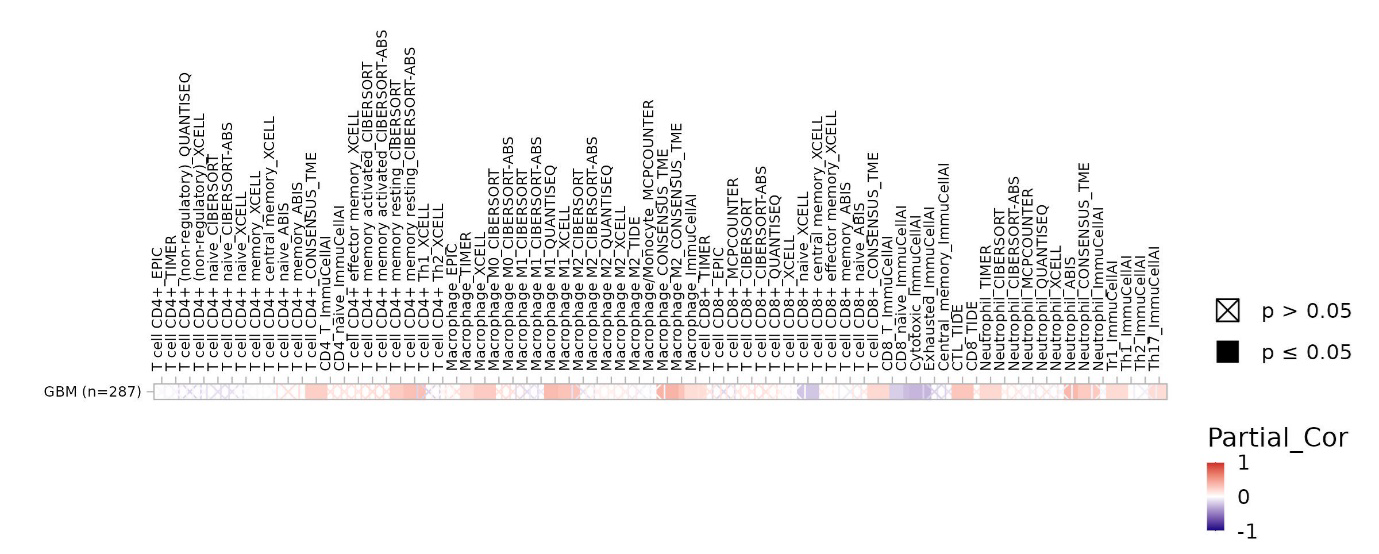
Figure 1. TIMER3 immune infiltration heatmap reveals modest associations between TACSTD2 expression and immune cell subsets in TCGA-GBM. Cells with a black square indicate p ≤0.05; cells with an X indicate p >0.05. CD4+ memory T-cell estimates (CIBERSORT, TME) show modest positive correlations (highlighted in the T-cell rows).
Purity-adjusted Spearman partial correlations (rho) between TACSTD2 expression and 50+ predicted immune cell populations in GBM (n=287 samples). Heatmap cells are colored by correlation strength: red = positive correlation (rho > 0), white = no correlation (rho ≈ 0), blue = negative correlation (rho <0). Black squares indicate statistically significant associations (p ≤0.05), while X-marked cells indicate p >0.05. Rows: individual immune cell subsets; columns: deconvolution algorithms and combined estimates. Most immune populations show weak correlations with TACSTD2 (rho <0.3), consistent with TACSTD2 not acting as a dominant immune checkpoint gene in GBM. However, CD4+ memory T cells across multiple algorithms show consistent modest positive correlations. CD8+ T cell estimates show predominantly near-zero correlations with no strong negative associations, indicating that TACSTD2-high tumors are not universally immune-excluded.
To visualize representative associations, we generated gene-specific scatter plots for TACSTD2 expression versus (i) CIBERSORT CD4+ memory resting T-cell estimates and (ii) CONSENSUS_TME CD4+ T-cell estimates in TCGA-GBM (Figure 2). In each plot, TACSTD2 expression (log2 TPM) is shown on the y-axis and immune infiltration level on the x-axis, with a less-smoothed trend line and 95% confidence interval. The corresponding Spearman rho and p-values reported by TIMER3 are displayed on the plots. A companion scatter of TACSTD2 versus tumor purity is shown for each algorithm to illustrate purity adjustment. All immune estimates derive from bulk RNA-seq deconvolution, which is known to be noisy in GBM due to extensive necrosis, variable tumor purity and myeloid-dominated infiltrates. We therefore interpret the TACSTD2-immune correlations as reflecting relative rather than absolute CD4+ memory abundance, that is, proportional enrichment of these subsets within the lymphocyte compartment rather than precise cell counts. Consistent with prior benchmarking of RNA-seq deconvolution methods in solid tumors, these algorithms tend to estimate relative rather than absolute immune cell fractions and are sensitive to tumor purity and stromal content. Benchmarking studies indicate that deconvolution methods are most reliable for relative cell-type fractions, particularly in complex tissues [11]. Our strategy of inferring cell-type associated programs from bulk RNA-seq and co-expression networks follows systems-biology approaches that have been successfully applied in other cancers [12].
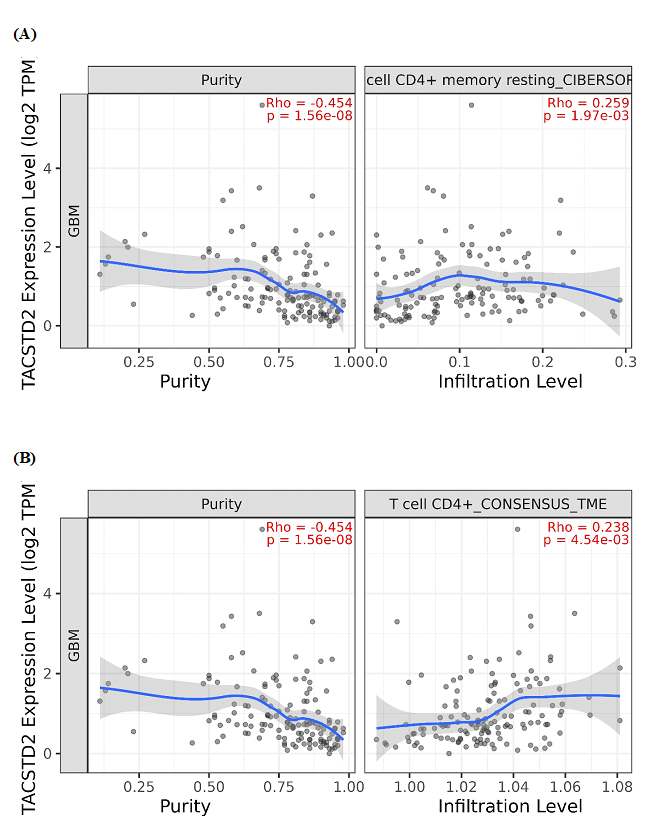
Figure 2. TACSTD2 expression positively correlates with CD4+ T-cell memory estimates across deconvolution algorithms. (A) Scatter plot of TACSTD2 expression (log2 TPM) versus CIBERSORT CD4+ memory resting T-cell infiltration in TCGA-GBM (n≈287). Each dot represents one tumor; the blue line shows a loess fit with 95% confidence interval (grey band). Spearman correlation rho and p-value are indicated in red. (B) Scatter plot of TACSTD2 expression (log2 TPM) versus T cell CD4+ CONSENSUS_TME infiltration estimates in the same cohort. A similar modest positive correlation (rho≈0.24, p≈4.5×10-3) supports reproducibility of the TACSTD2-CD4+ memory association across independent deconvolution methods.
Co-expression analysis and gene-set definition (GEPIA2)
To place TACSTD2 within co-expression networks, TACSTD2-correlated genes in GBM were identified using GEPIA2 (Correlation module):
- Gene: TACSTD2
- Cancer type: GBM (TCGA)
- Correlation method: Pearson
- Cut-offs: r >0.49, p <0.05
The top 100 positively correlated genes (highest Pearson r) were exported and used as the TACSTD2-associated gene set for pathway enrichment. This list was inspected to confirm enrichment for epithelial and junctional genes (e.g., claudins, cadherins, integrins, keratins).
Pathway enrichment analysis (Enrichr)
Pathway enrichment analysis of the TACSTD2-correlated gene set was performed using the Enrichr web server:
- Libraries interrogated: KEGG 2021 Human and MSigDB Hallmark 2020
- Statistics: Enrichr-reported p-values, Benjamini-Hochberg FDR-adjusted p-values and combined scores
Pathways with FDR <0.05 were considered statistically significant, but near-threshold immune-relevant pathways were also noted. For KEGG, enriched terms were visualized as a bar plot of the top pathways ranked by combined score (Figure 3A), highlighting hormone-related and adhesion pathways (e.g., estrogen signaling, ECM-receptor interaction, cell adhesion molecules, focal adhesion, adherens junction). For Hallmark gene sets, membership of TACSTD2-correlated genes in each hallmark was displayed using cluster-style heatmaps (Figures 3B and 3C), showing epithelial and junctional modules mapping to Estrogen Response (Early/Late) and Apical Junction signatures.
These analyses are based on co-expression and therefore infer biological programs indirectly; enrichment is interpreted as supportive of hormone-responsive epithelial and adhesion modules associated with TACSTD2, rather than as proof of pathway activation in every TACSTD2-high tumor.
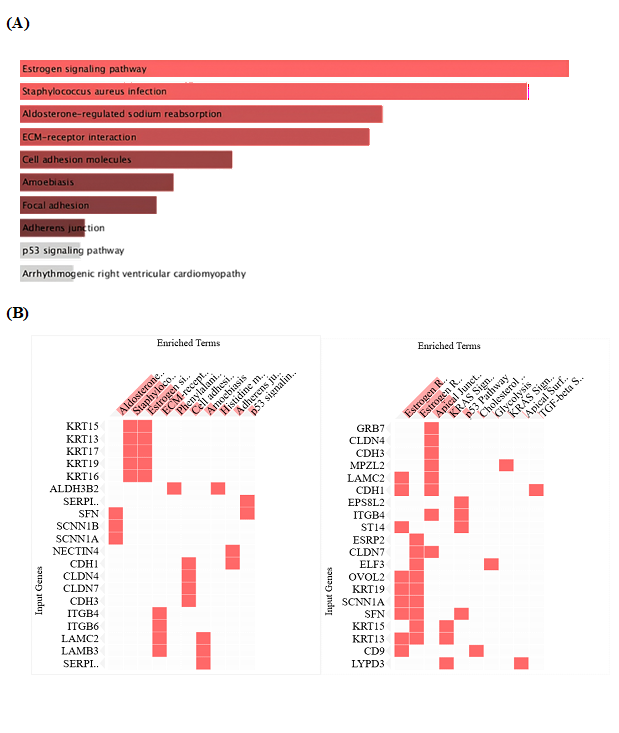
Figure 3. Pathway enrichment of TACSTD2-correlated genes reveals hormone-responsive epithelial programs. (A) KEGG 2021 Human pathway enrichment. Top KEGG 2021 Human pathways enriched among TACSTD2-correlated genes. Estrogen signaling is the top pathway, followed by adhesion-related pathways (ECM–receptor interaction, cell adhesion molecules, focal adhesion, adherens junction). Bar graph showing top enriched pathways ranked by combined score. Estrogen signaling pathway emerges as the top-ranked pathway (p=6.5×10-4, combined score ≈ 93), followed by aldosterone-regulated sodium reabsorption, and ECM-receptor interaction. Cell adhesion-related pathways (focal adhesion, adherens junction) also show significant enrichment. (B) MSigDB Hallmark 2020 cluster gram. Cluster gram showing TACSTD2-correlated genes (rows) versus enriched pathways (columns). Many epithelial/junction genes (e.g. CLDN4, CLDN7, CDH1, ITGB4, LAMC2) map to Estrogen Response and Apical Junction pathways, whereas few genes fall into classical immune checkpoint pathways. Heat map showing TACSTD2-correlated genes (rows) versus enriched Hallmark pathways (columns). Red squares indicate gene membership in pathway; white indicates absence. Estrogen Response Early and Late hallmarks show the strongest enrichment, alongside Apical Junction. Keratin family proteins (KRT15, KRT13, KRT19) cluster with Apical Junction signatures. Claudins (CLDN4, CLDN7), cadherins (CDH1, CDH3), and integrins (ITGB4, LAMC2) cluster with Estrogen Response hallmarks, reflecting hormone-tuned epithelial barrier control.
Results
TACSTD2 expression is aberrantly reactivated in a subset of GBM
Pan-cancer analysis of TCGA and GTEx datasets in GEPIA2 confirmed that TACSTD2 is highly expressed in multiple epithelial tumors but is nearly absent from most normal brain tissues. In the GBM cohort, TACSTD2 expression was detectable in a subset of tumors at levels substantially higher than normal brain, consistent with aberrant reactivation of an epithelial lineage marker in glial tissue.
TACSTD2 expression shows modest but reproducible association with CD4+ memory T-cell infiltration
TIMER3 Immune_Gene analysis generated a heatmap summarizing tumor-purity-adjusted Spearman partial correlations between TACSTD2 expression and inferred abundance of >50 immune cell subsets across multiple deconvolution algorithms in TCGA-GBM (n≈287; Figure 1). Most immune populations demonstrated weak correlations with TACSTD2 (rho < 0.3), consistent with TACSTD2 not acting as a dominant immune checkpoint gene in GBM. However, several CD4+ T-cell subsets showed reproducible positive associations. Deconvolution-based analyses of tumor immunity have reported similar correlation magnitudes (≈0.2-0.3) for chemokines such as CXCL9/10 with T-cell subsets [10,13,14].
The most consistent association was observed with CD4+ memory estimates. In the CIBERSORT algorithm, TACSTD2 expression correlated positively with CD4+ memory resting T-cell infiltration (rho≈0.26, p≈2×10³) after adjustment for tumor purity, with tumors expressing higher TACSTD2 tending to show higher inferred CD4+ memory levels (Figure 2A). A similar modest positive correlation was observed for the CONSENSUS_TME CD4+ T-cell estimate (rho≈0.24, p≈4.5×10-3; Figure 2B). These effect sizes are modest in absolute magnitude and are interpreted as a correlative trend. Nonetheless, they fall within the range reported for established chemokines and immune-modulatory genes in deconvolution-based analyses of tumor immunity, indicating that the TACSTD2-CD4+ memory association is not unusually weak for bulk RNA-seq deconvolution.CD4+ memory resting T-cell estimates were obtained from the CIBERSORT algorithm as implemented in TIMER3 [15]. Our analysis relies on deconvolution of bulk RNA-seq to infer immune cell subsets, an approach that has been widely applied to characterize tumor immune landscapes across cancer types [16].
In contrast, correlations between TACSTD2 expression and CD8+ T-cell estimates were close to zero across algorithms, and no strong negative association with global CD8+ infiltration was detected. Macrophage, neutrophil, and other myeloid estimates showed only heterogeneous, low-magnitude correlations. Together, these observations suggest that TACSTD2 reactivation in GBM is preferentially associated with relative enrichment of CD4+ memory T-cell populations rather than with a global increase or depletion of lymphoid or myeloid infiltrates.
TACSTD2 does not behave as a canonical immune-exclusion marker in GBM
In triple-negative breast cancer and other epithelial malignancies, TROP2 participates in a tight-junction/claudin program that enforces exclusion of CD8+ T cells from tumor nests and confers resistance to PD-1/PD-L1 blockade [17]. The TIMER3 analysis in GBM does not support a similar immune-exclusion model for TACSTD2. TACSTD2 expression lacked a strong negative correlation with overall CD8+ T-cell abundance, and the dominant association in GBM was instead a modest positive correlation with CD4+ memory T-cell estimates. This pattern indicates that TACSTD2-high GBM is not necessarily CD8-depleted and may coexist with preserved or relatively enriched adaptive immune presence, differing from the immune-cold phenotype described for TROP2-high epithelial tumors.
TACSTD2-correlated gene network highlights hormone-responsive epithelial junction programs
GEPIA2 correlation analysis identified 100 genes most strongly co-expressed with TACSTD2 in TCGA-GBM (r >0.49, p <0.05). The top positively correlated genes included multiple epithelial and junctional components such as keratins (KRT13, KRT15, KRT17, KRT19), claudins (CLDN4, CLDN7), cadherins (CDH1, CDH3) and integrins (ITGB4, ITGB6, LAMC2), forming a cohesive epithelial/junctional module.
Enrichr KEGG 2021 Human enrichment analysis of this gene set revealed estrogen signaling pathway as the top enriched pathway, followed by Staphylococcus aureus infection, aldosterone-regulated sodium reabsorption, ECM-receptor interaction, cell adhesion molecules, focal adhesion, and adherens junction (Figure 3A). MSigDB Hallmark 2020 analysis further emphasized hormone and junction biology: Estrogen Response Early, Estrogen Response Late and Apical Junction were among the most significantly enriched hallmarks (Figures 3B and 3C). Cluster-style heatmaps showed that claudins, cadherins, and integrins were strongly represented in estrogen-response and apical-junction signatures, consistent with a hormone-tuned epithelial barrier program.
Notably, TACSTD2-correlated genes showed minimal membership in canonical immune-checkpoint hallmark sets, suggesting that TACSTD2 marks a lineage and adhesion program rather than a classic T-cell-intrinsic suppressive checkpoint. When considered together with the deconvolution results, these enrichment patterns support a model in which TACSTD2-high GBM defines a hormone-responsive epithelial junction program that co-varies with relative enrichment of CD4+ memory T-cell infiltration, rather than a purely immune-excluded CD8-low state.
Discussion
This in silico analysis provides initial evidence that TACSTD2 expression in GBM is associated with specific features of the immune microenvironment and with hormone-responsive epithelial junction programs. Using TIMER3, TACSTD2 expression showed a reproducible, tumor-purity-adjusted positive correlation with CD4+ memory T-cell estimates across two independent deconvolution algorithms (CIBERSORT and CONSENSUS_TME), while lacking a strong negative association with global CD8+ T-cell abundance. Similar systems-biology frameworks integrating bulk expression, co-expression networks and pathway enrichment have been used to identify prognostic subnetworks in breast cancer supporting the use of such correlative network-based analyses as hypothesis-generating tools rather than definitive mechanistic proof [12].
Pathway analysis of TACSTD2-correlated genes highlighted estrogen signaling, estrogen-response, and apical junction gene sets, along with ECM-receptor interaction, focal adhesion and cell-adhesion molecules, with minimal overlap with canonical immune checkpoint pathways. Taken together, these modest but consistent associations support a model in which TACSTD2 marks a hormone-tuned epithelial junction program that co-varies with relative enrichment of CD4+ memory T cells, rather than functioning as a dominant immune-exclusion checkpoint in GBM.
TACSTD2 and adaptive immune context in GBM
The functional role of CD4+ memory T cells in GBM is complex and context-dependent. CD4+ memory and tissue-resident memory T cells can contribute to anti-tumor immunity and durable tumor control in glioblastoma models and patients, particularly in the context of virus-specific or neoantigen-specific responses.
The positive association between TACSTD2 expression and CD4+ memory T-cell estimates suggests that TACSTD2-high GBM is not simply immunologically inert. GBM-infiltrating T cells frequently exhibit severe exhaustion signatures, with impaired effector function [18]. CD4+ memory and tissue-resident memory T cells can contribute to anti-tumor immunity and durable tumor control in glioblastoma models and patients, particularly in the context of virus-specific or neoantige-specific responses, although their net impact on prognosis depends on functional state, localization, and interaction with myeloid and stromal compartments. At the same time, T-cell infiltrates in GBM frequently display exhaustion signatures and may include regulatory or Th2-skewed CD4+ subsets with immunosuppressive or tumor-supportive roles. Recent work has shown that virus-specific memory T cells can survey glioblastoma and mediate durable control in experimental settings [19,20].
Our bulk deconvolution analysis cannot distinguish between these functional phenotypes, and the correlation coefficients between TACSTD2 and CD4+ memory are modest (rho≈0.24-0.26), so the findings are best viewed as indicating a relative enrichment of CD4+ memory-like signatures in TACSTD2-high tumors rather than a definitive prognostic cell type. Nevertheless, the reproducibility of the TACSTD2-CD4+ memory association across two algorithms, together with prior evidence that T-cell abundance and exhaustion state influence GBM outcome and response to therapy, supports further investigation of how TACSTD2-defined epithelial programs intersect with adaptive immunity.
Divergence from epithelial TROP2 immune-exclusion models
In triple-negative breast cancer and other epithelial malignancies, TROP2 participates in a claudin-associated tight-junction program that enforces CD8+ T-cell exclusion from tumour nests and confers resistance to PD-1/PD-L1 blockade. If TACSTD2 played an analogous role in GBM, one would expect TACSTD2-high tumours to show strong negative correlations with CD8+ T-cell estimates and a more globally immune-cold phenotype. Instead, the deconvolution data show no strong negative association with CD8+ T-cell infiltration and a modest positive association with CD4+ memory T-cell signatures.
These differences likely reflect lineage- and context-specific functions of TACSTD2. GBM arises from glial or progenitor populations within the CNS, with distinct stromal architecture, myeloid composition, and barrier structures compared with epithelial tissues. Glioma growth is strongly influenced by myeloid-derived cells recruited by factors such as S100B, illustrating the importance of non-lymphoid components of the GBM microenvironment. Glioma growth is strongly influenced by myeloid-derived cells recruited by factors such as S100B, illustrating the importance of non-lymphoid components of the GBM microenvironment [21]. In this setting, TACSTD2 may integrate into an epithelial-like adhesion and hormone-response program without necessarily recreating the same physical barrier properties or immune-exclusion mechanisms observed in epithelial carcinomas. The data therefore favor a view of TACSTD2 as a marker of lineage and adhesion rewiring that correlates with a particular immune milieu, rather than as a directly targetable immune checkpoint in GBM.
Hormone-responsive epithelial junction program as an upstream axis
The enrichment of estrogen signaling, estrogen-response, and apical junction pathways among TACSTD2-correlated genes provides an interpretive link between TACSTD2 reactivation, sex-modulated methylation, and immune context in GBM. The TACSTD2-associated gene set includes multiple claudins, cadherins, integrins, and keratins, consistent with a partial acquisition of epithelial junctional identity in TACSTD2-high tumors. Previous work has shown sex-modulated CpG methylation at the TACSTD2 promoter and sex-stratified survival effects in GBM, raising the possibility that differential methylation gates access of hormone receptors (e.g., ESR1, AR, FOXA1) to the TACSTD2 locus in a sex-dependent manner.
Although the present analysis does not explicitly stratify by sex, the combined methylation, pathway, and deconvolution findings support a hypothesis in which sex-modulated TACSTD2 methylation regulates a hormone-responsive epithelial junction program that shapes, or at least co-varies with, the CD4+ memory T-cell landscape in TACSTD2-high GBM. Testing this model will require integration of sex-stratified methylation, hormone-receptor expression, immune infiltration, and spatial context at higher resolution [22].
Finally, while TACSTD2/TROP2-targeted antibody-drug conjugates are in development for epithelial cancers, the present data suggest that TACSTD2-high GBM may not share the same CD8+ T-cell exclusion phenotype. Any extrapolation of TROP2-directed strategies to GBM should therefore be considered speculative and contingent on preclinical studies that define how TACSTD2 modulation influences both tumor-intrinsic biology and CD4+ memory-rich immune landscapes in the brain.
Limitations
Several limitations should be emphasized. First, all immune infiltration estimates are derived from deconvolution of bulk RNA-seq data; in GBM, such estimates are affected by extensive necrosis, variable tumor purity, and myeloid-dominated infiltrates, and are more reliable for relative than absolute lymphocyte fractions. The TACSTD2-CD4+ memory correlations (rho≈0.24-0.26) are therefore interpreted as modest, hypothesis-generating trends rather than precise quantitative biomarkers of CD4+ memory cell density. Second, the analysis is cross-sectional and correlative; no causal inference can be made about whether TACSTD2 expression actively shapes immune infiltration or simply tracks with other underlying programs.
Third, the pathway enrichment results infer biological programs indirectly from co-expression and do not guarantee functional activation of estrogen signaling or junctional pathways in every TACSTD2-high tumor. Fourth, we were unable to perform robust sex-stratified immune deconvolution within TIMER3/GEPIA2 due to limited sex annotation and small subgroup sizes in TCGA-GBM, which is a major limitation given prior sex-modulated methylation and survival effects at the TACSTD2 locus. Finally, this work focuses on TACSTD2 as a single-gene entry point; more comprehensive multi-gene or network-level approaches may capture additional immune-relevant axes that are not apparent from TACSTD2 alone.
Future Directions
Future work should integrate these in silico observations with single-cell RNA-seq and spatial transcriptomic datasets of GBM to map TACSTD2 expression at cellular resolution and delineate its relationship to specific CD4+ memory and tissue-resident subsets, as well as to exhausted and regulatory phenotypes. Spatial transcriptomics has revealed segregation of tumor cell states and marked immunosuppression within peri-necrotic niches in GBM [23]. In addition, spatial profiling studies have shown that GBM harbors distinct tumor cell states and immune niches such as peri-necrotic regions with marked immunosuppression which could help localize TACSTD2-high epithelial-like programs and their associated immune communities.Integrative spatial analysis, thus, further demonstrated the multilayered organization of GBM and its immune microenvironments. Integrative spatial analysis has further demonstrated the multilayered organization of GBM and its immune microenvironments [18,24].
Sex-stratified analyses of immune infiltration in TACSTD2-high versus TACSTD2-low tumors, combined with TACSTD2 promoter methylation and hormone-receptor expression data, will be important for understanding how sex-biased epigenetic regulation intersects with immune context in GBM [25,26]. In parallel, functional studies that manipulate TACSTD2 expression in GBM models and assess effects on junctional architecture, myeloid recruitment and CD4+ memory T-cell behavior will be required to determine whether TACSTD2 is merely a marker of an epithelial-hormone program or an active participant in shaping the tumor microenvironment.
Finally, while TACSTD2/TROP2-targeted antibody-drug conjugates are in development for epithelial cancers, the present data suggest that TACSTD2-high GBM may not share the same CD8+ T-cell exclusion phenotype. Any extrapolation of TROP2-directed strategies to GBM should therefore be considered speculative and contingent on preclinical studies that define how TACSTD2 modulation influences both tumor-intrinsic biology and CD4+ memory-rich immune landscapes in the brain.
Conclusions
This computational study identifies TACSTD2 as a marker of a hormone-responsive epithelial junction program that co-varies with modest enrichment of CD4+ memory T-cell signatures in GBM, without evidence of strong CD8+ T-cell exclusion. The findings are correlative and derived from bulk RNA-seq deconvolution, but they provide a testable framework linking sex-modulated TACSTD2 reactivation, epithelial lineage rewiring, and adaptive immune context that can guide future single-cell, spatial, and functional investigations in glioblastoma [27].
Data Availability
All data analysed in this study are publicly available from TCGA and associated portals accessed through TIMER3, GEPIA2, and Enrichr; no new experimental datasets were generated.
Author Contributions
T.F. conceived the study, performed all analyses, interpreted the data and wrote the manuscript.
Funding
This work received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Conflicts of Interest
The author declares no competing interests.
References
2. Huang R, Lu X, Sun X, Wu H. A novel immune cell signature for predicting glioblastoma after radiotherapy prognosis and guiding therapy. Int J Immunopathol Pharmacol. 2024 Jan-Dec;38:3946320241249395.
3. Zhang W, Zhang W, Wu H, Han X. Harnessing innate immunity against glioblastoma microenvironment. Front Immunol. 2025 Jul 25;16:1648601.
4. Wu B, Thant W, Bitman E, Liu T, Liu J, Paschalis EI, et al. A TROP2/Claudin Program Mediates Immune Exclusion to Impede Checkpoint Blockade in Breast Cancer. bioRxiv [Preprint]. 2024 Dec 5:2024.12.02.626446.
5. Liu X, Deng J, Yuan Y, Chen W, Sun W, Wang Y, et al. Advances in Trop2-targeted therapy: Novel agents and opportunities beyond breast cancer. Pharmacol Ther. 2022 Nov;239:108296.
6. Hou J, Lv A, Deng Q, Zhang G, Hu X, Cui H. TROP2 promotes the proliferation and metastasis of glioblastoma cells by activating the JAK2/STAT3 signaling pathway. Oncol Rep. 2019 Feb;41(2):753–64.
7. Zaman S, Jadid H, Denson AC, Gray JE. Targeting Trop-2 in solid tumors: future prospects. Onco Targets Ther. 2019 Mar 1;12:1781–90.
8. Shaffer C. Trop2 deal heats up antibody-drug conjugate space in cancer. Nat Biotechnol. 2021 Feb;39(2):128–30.
9. Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, et al. TIMER: A Web Server for Comprehensive Analysis of Tumor-Infiltrating Immune Cells. Cancer Res. 2017 Nov 1;77(21):e108–10.
10. Li B, Severson E, Pignon JC, Zhao H, Li T, Novak J, et al. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol. 2016 Aug 22;17(1):174.
11. Hu M, Chikina M. Heterogeneous pseudobulk simulation enables realistic benchmarking of cell-type deconvolution methods. Genome Biol. 2024 Jul 1;25(1):169.
12. Song Q, Wang H, Bao J, Pullikuth AK, Li KC, Miller LD, et al. Systems biology approach to studying proliferation-dependent prognostic subnetworks in breast cancer. Sci Rep. 2015 Aug 10;5:12981.
13. La Manna MP, Di Liberto D, Lo Pizzo M, Mohammadnezhad L, Shekarkar Azgomi M, Salamone V, et al. The Abundance of Tumor-Infiltrating CD8+ Tissue Resident Memory T Lymphocytes Correlates with Patient Survival in Glioblastoma. Biomedicines. 2022 Oct 1;10(10):2454.
14. Ning J, Gavil NV, Wu S, Wijeyesinghe S, Weyu E, Ma J, et al. Functional virus-specific memory T cells survey glioblastoma. Cancer Immunol Immunother. 2022 Aug;71(8):1863–75.
15. Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015 May;12(5):453–7.
16. Luo J, Liu L, Venkateswaran S, Song Q, Zhou X. RPI-Bind: a structure-based method for accurate identification of RNA-protein binding sites. Sci Rep. 2017 Apr 4;7(1):614.
17. Ambrogi F, Fornili M, Boracchi P, Trerotola M, Relli V, Simeone P, et al. Trop-2 is a determinant of breast cancer survival. PLoS One. 2014 May 13;9(5):e96993.
18. Woroniecka K, Chongsathidkiet P, Rhodin K, Kemeny H, Dechant C, Farber SH, et al. T-Cell Exhaustion Signatures Vary with Tumor Type and Are Severe in Glioblastoma. Clin Cancer Res. 2018 Sep 1;24(17):4175–86.
19. Ning J, Gavil NV, Wu S, Wijeyesinghe S, Weyu E, Ma J, et al. Correction to: Functional virus specific memory T cells survey glioblastoma. Cancer Immunol Immunother. 2022 Oct;71(10):2577–79.
20. Ning J, Gavil NV, Wu S, Wijeyesinghe S, Weyu E, Ma J, et al. Functional virus-specific memory T cells survey glioblastoma. Cancer Immunol Immunother. 2022 Aug;71(8):1863–75.
21. Wang H, Zhang L, Zhang IY, Chen X, Da Fonseca A, Wu S, et al. S100B promotes glioma growth through chemoattraction of myeloid-derived macrophages. Clin Cancer Res. 2013 Jul 15;19(14):3764–75.
22. Liu M, Ji Z, Jain V, Smith VL, Hocke E, Patel AP, et al. Spatial transcriptomics reveals segregation of tumor cell states in glioblastoma and marked immunosuppression within the perinecrotic niche. Acta Neuropathol Commun. 2024 Apr 22;12(1):64.
23. Ravikumar V, Maddox A, Kulkarni R, Rao A, Al-Holou WN. Spatial transcriptomics of glioblastoma defines biologically and clinically significant reprogramming patterns across unique spatial microenvironments. bioRxiv [Preprint]. 2025 Oct 26:2025.10.17.683110.
24. Greenwald AC, Darnell NG, Hoefflin R, Simkin D, Mount CW, Gonzalez Castro LN, et al. Integrative spatial analysis reveals a multi-layered organization of glioblastoma. Cell. 2024 May 9;187(10):2485–2501.e26.
25. Stepan LP, Trueblood ES, Hale K, Babcook J, Borges L, Sutherland CL. Expression of Trop2 cell surface glycoprotein in normal and tumor tissues: potential implications as a cancer therapeutic target. J Histochem Cytochem. 2011 Jul;59(7):701–10.
26. Trerotola M, Cantanelli P, Guerra E, Tripaldi R, Aloisi AL, Bonasera V, et al. Upregulation of Trop-2 quantitatively stimulates human cancer growth. Oncogene. 2013 Jan 10;32(2):222–33.
27. Lang X, Zhang S, Wang Y. Targeting the neuroimmune axis in glioblastoma: emerging strategies for precision immunotherapy. Front Immunol. 2025 Sep 9;16:1661327.
