Abstract
Cell adhesion molecules (CAM) mediate cell to cell interactions in various body systems including the immune system. The four major families of CAMs include immunoglobulin (Ig) superfamily, cadherins, integrins, and selectins. Most interestingly, several CAMs have emerged in recent years as leading biomarkers of lupus nephritis (LN) based on comprehensive proteomic screens of urine. Proteins belonging to all four families of CAMs have been reported to be upregulated in LN urine. Belonging to the Ig superfamily are the following elevated proteins: ALCAM, VCAM-1, BCAM, EpCAM, NCAM-1, NCAM-120, NrCAM, OBCAM, PECAM-1, sICAM-1, sICAM-2, and sICAM-5. Similarly, cadherins such as P-Cadherin, Cadherin-5, Cadherin-2, Cadherin-12, and E-Cadherin, integrin family members integrin avb5, integrin a1b1, fibronectin, and GP1BA, as well as multiple selectins including L-selectin, E-selectin, and P-selectin have all been reported to be upregulated in LN urine. These findings and the documented interactions between these proteins and their ligands suggest that “sticky interactions” between various immune cells mediated by CAMS may play key pathogenic roles in this disease.
Commentary
Cell adhesion molecules (CAM) mediate cell to cell interactions in various body systems including the immune system. The four major families of CAMs include immunoglobulin (Ig) superfamily, cadherins, integrins, and selectins. Most interestingly, several CAMs have emerged in recent years as leading biomarkers of lupus nephritis (LN), based on comprehensive proteomic screens of urine. These findings suggest that “sticky interactions” between various immune cells mediated by CAMS may play key pathogenic roles in this disease.
The activated leukocyte cell adhesion molecule (ALCAM) is a surface transmembrane glycoprotein from the immunoglobulin superfamily. The protein is expressed on antigen presenting cells (APCs) including macrophages, B cells, and more particularly on dendritic cells (DCs). ALCAM, also known as CD166, serves as a CD6 ligand, co-stimulating T cells and enabling immune cell adhesion through CD6 binding or through ALCAM-ALCAM interactions. In a recent study, Ding et al. showed that urinary ALCAM may serve as a potential biomarker to differentiate active LN from inactive systemic lupus erythematosus (SLE), SLE without renal involvement, and healthy control subjects in a Chinese cohort [1]. Elevated urinary ALCAM levels in LN patients significantly correlated with clinical parameters - renal SLEDAI and SLICC - as well as other laboratory parameters such as 24hr urine, serum complement C3, and hemoglobin. Urinary ALCAM distinctly outperformed conventional LN markers including serum dsDNA, C3, C4, and proteinuria. Hence, urinary ALCAM may serve as a better diagnostic biomarker for predicting renal pathology as well as an alternative diagnostic marker for renal histology in LN. The findings of the study revealed that urinary ALCAM was expressed most significantly in active LN patients compared to the remaining three groups. In contrast to ALCAM, traditional biomarkers including anti-dsDNA and anti-C1q antibodies, hypocomplementaemia and albuminuria were unable to reliably differentiate between class III/IV and V lupus nephritis and accurately predict long-term renal function impairment [1].
This study [1] joins a number of articles establishing ALCAM as a novel biomarker of LN histopathology. Collectively, these studies reveal that ALCAM is a reliable urinary biomarker of LN in multiple ethnic groups, including African American, Hispanic, Caucasian, and Chinese [2]. Within these four cohorts, greater levels of ALCAM present in the urine of LN patients correlated with increased disease severity and renal involvement. These findings once again point to the potential pathogenic relevance of ALCAM-CD6 interactions. The ALCAM-CD6 pathway has been previously studied for its involvement in the pathogenesis of several autoimmune conditions including rheumatoid arthritis, psoriasis, multiple sclerosis, IBD, as well as specific cancers [3-7]. The abundance of clinical data and animal studies underscore the crucial contributions the ALCAM-CD6 receptor-ligand complex makes in autoimmune disease progression. Based on its known function, ALCAM is hypothesized to contribute to the pathogenesis of LN through its interaction with CD6 in a number of ways.
The pathogenic role of T cells in LN is well established by animal and clinical studies as prime regulators of kidney inflammation [8,9]. Found on the surface of T cells is CD6, a transmembrane glycoprotein. When CD6 associates with its ligand ALCAM, it co-stimulates T-cell activation and proliferation [10]. As a crucial mediator of T cell activation, ALCAM-CD6 complex localizes at the DC-T cell interface and may induce the differentiation of multiple T helper (Th) cells, including Th1, Th2, Th17 and CD8 T cell subsets [11-12]. This ability is evidenced by a study in which ALCAM-CD6 interaction was blocked through anti-CD6 dosing, preventing normal differentiation of CD4 T cells [13]. Moreover, recent in vitro and in vivo studies blocked ALCAM-CD6 interaction in patients with active rheumatoid arthritis (RA) and psoriasis patients using an anti-CD6 monoclonal antibody [14,15]. Clinical benefits were reported in both studies: RA patients experienced reduced and alleviated adverse events and psoriasis patients noted substantial amelioration of epidermis hyperplasia [14,15].
CD4 and CD8 T cells are prominent renal-infiltrating immune cells that contribute to the pathogenesis of LN by promoting disease progression through the regulation and activation of B cells, myeloid cells and other intra-renal cells [16]. Greater numbers of CD4+ and CD8+ T cells have been documented within the kidneys of lupus-prone mice as well as the kidneys of patients with severe LN [14,15]. CD6 expression on renal-infiltrating T cells juxtaposed to ALCAM expressing antigen presenting cells may allow for CD6-ALCAM interactions. CD6 and ALCAM bind and colocalize with TCR/ CD3 at the immunological synapse, effectively stimulating T-cell activation, differentiation and proliferation [17-19]. Unpublished data from our lab shows that specific immune cell types that are key regulators of lupus pathogenesis highly express CD6 and ALCAM in lupus mice models, demonstrating that CD6-ALCAM signaling could further induce renal CD4 and CD8 T cell recruitment and activation, contributing to tissue inflammation. Together, these combined mechanisms sustain increased T cell activation and trafficking to maintain progressive renal injury, potentially driving the pathogenesis of LN.
Beyond its role as an exceptional biomarker for LN, recent studies attribute a pathogenic role for ALCAM in LN. In a recently reported abstract, the ALCAM-CD6 pathway was pharmacologically blocked using anti-CD6 monoclonal antibodies, resulting in decreased levels of renal-infiltrating T cells and subsequent decrease in the severity of renal disease and LN in mice [20]. This was accompanied by significant improvement in the manifestations of lupus and LN, notably lymphadenopathy, skin lesions, and renal disease [20]. Hence, ALCAM is not only an established biomarker for LN but also a pathogenic driver. Currently underway are clinical and mechanistic studies exploring the precise manner in which ALCAM-CD6 pathway blockade leads to reduced kidney inflammation in LN.
ALCAM represents one of many cell adhesion molecules (CAMs) found to be elevated in the urine of active LN patients. As exemplified by ALCAM, CAMs play an integral role in facilitating inflammatory processes primarily by allowing adhesion of leukocytes to the vascular endothelium as well as transendothelial migration at sites of inflammation [21]. The four major families of CAMs include immunoglobulin (Ig) superfamily, cadherins, integrins, and selectins.
As stated above, ALCAM belongs to the immunoglobulin (Ig) superfamily. Urinary ALCAM is observed to be significantly increased in active LN, with a 14-fold change (FC) in LN compared to healthy control, (p=0.02) [2]. In addition to ALCAM, within the Ig superfamily is vascular CAM1 (VCAM-1), a prominent CAM ~ 8.9-fold higher compared to healthy control (p=0.0003) [2]. Other proteins within the Ig superfamily that are elevated compared to control include BCAM (FC=3.7, p=0.02), EpCAM (FC=35.7, p=0.02), NCAM-1 (FC=10.7, p=0.002), NCAM- 120 (FC=6.4, p ≤ 0.009), NrCAM (FC=2.5, p=0.01), OBCAM (FC=5.6, p=0.0003), PECAM-1 (FC=2.6, p=0.006), sICAM-1 (FC=6.2, p=0.0003), sICAM-2 (FC=2.8, p=0.001), and sICAM-5 (FC=2.8, p=0.001) [2,22], as listed in Table 1.
| CAM Family Biomarker Protein | Potential Ligands * | Fold Change1 | P value1 |
|---|---|---|---|
| GP1BA | VWF | 4.5 | 0.0012 |
| Integrins Integrin a1b1 | Collagens I, IV, VI, XIII, XVI, laminin | 12.6 | 0.0205 |
| Integrin aVb5 | vitronectin, fibronectin, MMP-2, OPN, laminin, VWF | 6.5 | 0.0006 |
| Cadherin-12 | other cadherins | 2.4 | 0.0093 |
| Cadherin-2 | CDH11, CTNNA1, CTNND1, B-catenin, LRRC7, PTPRM | 4.2 | 0.0022 |
| Cadherins Cadherin-5 | B-catenin, plakoglobin, PTPRB, CTNND1 | 47.0 | 0.0003 |
| E-Cadherin | B-catenin, | 28.2 | 0.0040 |
| P-Cadherin | CDH1, B-catenin, nephrin, plakoglobin | 3.7 | 0.0003 |
| E-Selectin | CD44, PSGL-1, LAMP1, LAMP2, Mac2-BP | 13.2 | 0.0006 |
| Selectins L-Selectin | GlyCAM-1, CD34, MadCAM-1, PSGL-1 | 5.8 | 0.0003 |
| P-Selectin | PSGL-1 | 5.1 | 0.0059 |
| ALCAM | CD6 | 18.1 | 0.0205 |
| BCAM | Laminin | 3.7 | 0.0205 |
| EpCAM | CLDN7, ACTN1 | 35.7 | 0.0186 |
| ICAM-1 | CD11/CD18, fibrinogen | 6.2 | 0.0003 |
| ICAM-2 | EZR, P9, CD11/CD18 | 8.1 | 0.0001 |
| ICAM-5 | PSEN1 | 2.8 | 0.0012 |
| Ig superfamily NCAM-1 | NCAM-1 | 10.7 | 0.0016 |
| NCAM-120 | NCAM-1 | 6.4 | 0.0093 |
| Nr-CAM | Nrp-2 | 2.5 | 0.0140 |
| OBCAM | Opiod ligands | 5.6 | 0.0003 |
| PECAM-1 | CD38, CD177, Integrin aVb3 | 2.6 | 0.0059 |
| VCAM-1 | VLA-4 | 25.2 | 0.0003 |
| * Listed are potential ligands of elevated CAM biomraker proteins in LN. If the ligand is also elevated in LN urine, it is bolded. 1: Indicated are the fold-change (and p-value) increase of biomarker protein in LN urine, as reported (2, 22). |
|||
Cadherins are expressed by epithelial cells and immune cells such as Langherhan’s cells, CD4+ and CD8+ T cells, murine cutaneous vγ5+γ/δ T cells, erythrocytes and normoblasts [23-26]. Cadherins shows hemophilic binding interactions and participate in cell recognition, tissue morphogenesis and tumor suppression. On the cytoplasmic side, cadherins binds to catenins [27]. Cadherins elevated in LN include P-Cadherin (FC=3.7, p=0.0003), Cadherin-5 (FC=47.0, p=0.0003), Cadherin-2 (FC=4.2, p=0.002), Cadherin-12 (FC=2.4, p=0.009), and E-Cadherin (FC=28.2, p=0.004) [2,22], as listed in Table 1.
Integrins are expressed on various leukocytes and platelets [28]. GPIBa is a platelet surface glycoprotein composed of an alpha chain and a beta chain, that interacts with von Willebrand factor (VWF). This interaction facilitates platelet adhesion to the endothelium leading to enhanced platelet activation, thrombosis, and hemostasis. Integrin a1b1 (VLA1) is expressed predominantly on T-cells, and interacts with multiple collagens and laminin, widely expressed in multiple tissues and organs (Table 1). Integrin alpha V beta 5 is expressed on a wide variety of cell types including myeloid cells, keratinocytes, fibroblasts, endothelial and epithelial cells, where they bind ligands containing an RGD motif, including vitronectin, fibronectin, as well as MMP-2 and osteopontin, many of which are themselves elevated in LN urine (Table 1) [2,22].
The selectin family of proteins, expressed by several bonemarrow derived cells and endothelial cells [29-31], are also elevated in active LN urine compared to healthy control urine, including L-selectin (FC=5.8, p=0.0003), E-selectin (FC=13.2, p=0.0006), and P-selectin (FC=5.1, p=0.006) [2,22]. Leukocytes express L-selectins; endothelial cells express E-selectins; and platelets and endothelial cells express P-selectins. P-selectin glycoprotein ligand-1 (PSGL-1) is a major ligand for P-selectins. L-selectin ligands include GlyCAM-1, CD34, MadCAM-1, PSGL- 1 and heparan sulphate, while E-Selectin ligands include CD44, PSGL-1, LAMP1, LAMP2, Mac2-BP, as summarized in Table 1. In general, the ligands for the elevated selectins in LN do not themselves appear to be upregulated in LN urine (Table 1).
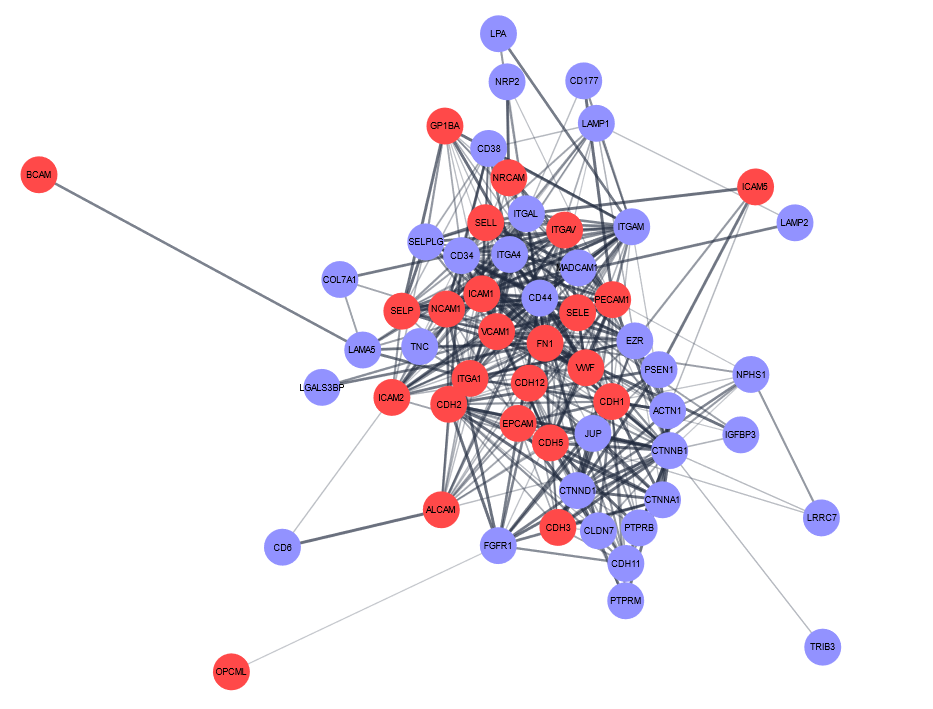
Interestingly, the CAMs and their respective ligands that were elevated in LN urine exhibited a rich pattern of interconnectedness (based on interactions with each other) as plotted using Cytoscape (Figure 1). The Reactome pathways enriched among the upregulated proteins in lupus nephritis included “Cell surface interaction, extracellular matrix organization, adherens junctions interactions, cell-cell communication, cell junction organization, and hemostasis”. Given the documented importance of CAMs within the inflammatory milieu in various pathogenic processes, the elevated CAM proteins elevated in LN are of importance and interest not only as biomarkers for LN, but also as potential drivers of disease pathogenesis, thus warranting further mechanistic studies. Unraveling the intricacies of these sticky interactions in LN may shed light on additional regulatory mechanisms contributing to LN disease progression and insight into further therapeutic opportunities in this complex systemic autoimmune disease.
References
2. Stanley S, Vanarsa K, Soliman S, Habazi D, Pedroza C, Gidley G, et al. Comprehensive aptamer-based screening identifies a spectrum of urinary biomarkers of lupus nephritis across ethnicities. Nature communications. 2020 May 4;11(1):1-3.
3. Alonso-Ramirez R, Loisel S, Buors C, Pers JO, Montero E, Youinou P, et al. Rationale for targeting CD6 as a treatment for autoimmune diseases. Arthritis. 2010;2010.
4. Montero E, Reyes G, Guibert M, Torres O, Rodriguez N, Estrada J, Torres L, Perez R, Hernandez A, Lage A. Immunodiagnosis and therapeutic immunosuppression in rheumatoid arthritis with ior t1 (anti-CD6) monoclonal antibody. InArthritis Research & Therapy 2002 Feb 4; 1: 1-38). BioMed Central.
5. Montero E, Falcon L, Morera Y, Delgado J, Amador JF, Perez R. CD6 molecule may be important in the pathological mechanisms of lymphocytes adhesion to human skin in psoriasis and ior t1 MAb a possible new approach to treat this disease. Autoimmunity. 1999 Jan 1;29(2):155-6.
6. Wagner M, Bilinska M, Pokryszko-Dragan A, Sobczynski M, Cyrul M, Kusnierczyk P, et al. ALCAM and CD6—multiple sclerosis risk factors. Journal of Neuroimmunology. 2014 Nov 15;276(1-2):98-103.
7. Ma C, Wu W, Lin R, Ge Y, Zhang C, Sun S, et al. Critical role of CD6highCD4+ T cells in driving Th1/Th17 cell immune responses and mucosal inflammation in IBD. Journal of Crohn’s and Colitis. 2019 Mar 30;13(4):510-24.
8. Okamoto A, Fujio K, Tsuno NH, Takahashi K, Yamamoto K. Kidneyinfiltrating CD4+ T-cell clones promote nephritis in lupus-prone mice. Kidney International. 2012 Nov 1;82(9):969-79.
9. Couzi L, Merville P, Deminière C, Moreau JF, Combe C, Pellegrin JL, et al. Predominance of CD8+ T lymphocytes among periglomerular infiltrating cells and link to the prognosis of class III and class IV lupus nephritis. Arthritis & Rheumatism: Official Journal of the American College of Rheumatology. 2007 Jul;56(7):2362-70
10. Jordan MA, Field J, Butzkueven H, Baxter AG. Genetic predisposition, humans. InThe Autoimmune Diseases 2014 Jan 1 (pp. 341-364). Academic Press.
11. Zimmerman AW, Joosten B, Torensma R, Parnes JR, van Leeuwen FN, Figdor CG. Long-term engagement of CD6 and ALCAM is essential for T-cell proliferation induced by dendritic cells. Blood. 2006 Apr 15;107(8):3212-20.
12. Bughani U, Saha A, Kuriakose A, Nair R, Sadashivarao RB, Venkataraman R, et al. T cell activation and differentiation is modulated by a CD6 domain 1 antibody Itolizumab. PloS one. 2017 Jul 3;12(7):e0180088.
13. Freitas RF, Basto A, Almeida SC, Santos RF, Gonçalves CM, Corria- Osorio J, et al. Modulation of CD4 T cell function via CD6-targeting. EBioMedicine. 2019 Sep 1;47:427-35.
14. Rodriguez PC, Torres-Moya R, Reyes G, Molinero C, Prada D, Lopez AM, et al. A clinical exploratory study with itolizumab, an anti- CD6 monoclonal antibody, in patients with rheumatoid arthritis. Results in Immunology. 2012 Jan 1;2:204-11.
15. Aira LE, López-Requena A, Fuentes D, Sánchez L, Pérez T, Urquiza A, Bautista H, et al. Immunological and histological evaluation of clinical samples from psoriasis patients treated with anti-CD6 itolizumab. InMAbs 2014 May 1; 6(3):782-92.
16. Suárez-Fueyo A, Bradley SJ, Tsokos GC. T cells in systemic lupus erythematosus. Current Opinion in Immunology. 2016 Dec 1;43:32-8.
17. Gimferrer I, Calvo M, Mittelbrunn M, Farnós M, Sarrias MR, Enrich C, et al. Relevance of CD6-mediated interactions in T cell activation and proliferation. The Journal of Immunology. 2004 Aug 15;173(4):2262-70.
18. Wagner M, Bilinska M, Pokryszko-Dragan A, Sobczynski M, Cyrul M, Kusnierczyk P, et al. ALCAM and CD6—multiple sclerosis risk factors. Journal of Neuroimmunology. 2014 Nov 15;276(1-2):98-103.
19. Sharp LL, Schwarz DA, Bott CM, Marshall CJ, Hedrick SM. The influence of the MAPK pathway on T cell lineage commitment. Immunity. 1997 Nov 1;7(5):609-18.
20. Chalmers S, Garcia S, Ramachandran RA, Mohan C, Ampudia J, Ng C, Connelly S, Putterman C. CD6 Modulation Ameliorates Skin and Kidney Disease in a Spontaneous Murine Model of SLE. Inarthritis & Rheumatology 2019 Oct 1; 71.
21. Meager A. Cytokine regulation of cellular adhesion molecule expression in inflammation. Cytokine & growth factor reviews. 1999 Mar 1;10(1):27-39.
22. Vanarsa K, Soomro S, Zhang T, Strachan B, Pedroza C, Nidhi M, et al. Quantitative planar array screen of 1000 proteins uncovers novel urinary protein biomarkers of lupus nephritis. Annals of the Rheumatic Diseases. 2020 Oct 1;79(10):1349-61.
23. Aiba S, Nakagawa S, Ozawa H, Tagami H. Different Expression of E-Cadherin by Two Cutaneous γ/δ TcR+ T-Cell Subsets, Vγ5-and Vγ5+ γ/δ TcR+ T cells. Journal of Investigative Dermatology. 1995 Sep 1;105(3):379-82.
24. Armeanu S, Bühring HJ, Reuss-Borst M, Müller CA, Klein G. E-cadherin is functionally involved in the maturation of the erythroid lineage. Journal of Cell Biology. 1995 Oct 1;131(1):243-9.
25. Caux C, Vanbervliet B, Massacrier C, Dezutter-Dambuyant C, de Saint-Vis B, Jacquet C, et al. CD34+ hematopoietic progenitors from human cord blood differentiate along two independent dendritic cell pathways in response to GM-CSF+ TNF alpha. The Journal of Experimental Medicine. 1996 Aug 1;184(2):695-706.
26. Munro SB, Duclos AJ, Jackson AR, Baines MG, Blaschuk OW. Characterization of cadherins expressed by murine thymocytes. Cellular Immunology. 1996 May 1;169(2):309-12.
27. Shapiro L, Weis WI. Structure and biochemistry of cadherins and catenins. Cold Spring Harbor perspectives in biology. 2009 Sep 1;1(3):a003053.
28. Luo BH, Carman CV, Springer TA. Structural basis of integrin regulation and signaling. Annual Review of Immunology. 2007 Apr 23;25:619-47.
29. Borsig L. Selectins in cancer immunity. Glycobiology. 2018 Sep;28(9):648-55.
30. Ley K, Kansas GS. Selectins in T-cell recruitment to non-lymphoid tissues and sites of inflammation. Nature Reviews Immunology. 2004 May;4(5):325-36.
31. Rosen SD. Ligands for L-selectin: homing, inflammation, and beyond. Annu. Rev. Immunol.. 2004 Apr 23;22:129-56.
