Abstract
For several decades, the transcription factor MEF2 (myocyte enhancer factor-2) has been known as a master regulator of myogenesis that orchestrates the first step in muscle formation: the differentiation of myoblasts into myocytes. Because of its importance during myoblast differentiation, the potential roles of MEF2 during later steps of myogenesis, in particular myocyte fusion, could not be properly investigated. Our recent findings filled that gap as we recently unravelled a key role of MEF2 in myocyte fusion both in vitro and in vivo. Mechanistically, we identified the PP2A-Bδ phosphatase as a key enzyme regulating MEF2 activity specifically during fusion, but not differentiation. We showed that by controlling phosphorylation of HDAC4, a known repressor of MEF2, PP2A-Bδ regulates the transcription of a subset of MEF2-dependent genes involved solely in myocyte fusion. Our observations demonstrate that, in contrast to the historical paradigm, MEF2 controls both early (i.e., the myoblast differentiation) and late (i.e., the myocyte fusion) steps of myogenesis with PP2A-Bδ allowing discrimination between both functions. Because PP2A holoenzymes exhibit high substrate specificity, our findings point towards novel therapeutic options for various myopathies characterized by defects in normal fusion, such as muscular dystrophies.
Introduction
Skeletal myogenesis is a highly complex and ordered process, entailing two main events [1,2]: (i) the specification of muscle precursors into myoblasts, followed by their differentiation into myocytes and (ii) the fusion of these differentiated myocytes giving rise to the multinucleated skeletal myofiber.
The class IIa Histone DeACetylases (HDACs)/MEF2 (Myocyte Enhancer Factor-2) axis has long being considered as the master regulator of the differentiation process during skeletal myogenesis [3]. However, our recent work [4] demonstrates that this pathway is also crucial for fusion of myocytes into myofibers, both in vitro and in vivo. Our findings thus further confirm and expend the idea that the class IIa HDACs/MEF2 axis is a cornerstone of the myogenic process in vertebrates
MEF2, the Historical Master Regulator of Myoblast Differentiation
Myoblast differentiation is coordinated by a hierarchical network of transcription factors, among which members of the MEF2 family, which includes MEF2a, -b, -c and -d play dominant roles [1,11]. MEF2 regulates the expression of a series of muscle structural genes and basic helixloop- helix (bHLH) transcription factors involved in the differentiation program, that are collectively known as myogenic regulatory factors (MRFs) and include Myf5, MyoD, myogenin and MRF4 [1]. The transcriptional activity of MEF2 is controlled by class IIa HDACs (HDAC4, -5, -7, and -9) and by MITR, a HDAC9 paralog lacking a HDAC catalytic domain [6-8]. Interestingly, the ability of class IIa HDACs to bind to MEF2 is determined by the phosphorylation status of conserved serine residues located in their amino-terminal non-HDAC domain [9,10]. When not phosphorylated on these residues, class IIa HDACs bind to MEF2 and repress its transcriptional activity. In contrast, phosphorylation of class IIa HDACs induces their detachment from MEF2, which restores the activity of the transcription factor [11].
At the beginning of the 2000’s, it mainly was the founding work of the Olson laboratory that brought to light the central role of the class IIa HDACs/MEF2 pathway in skeletal muscle development and remodeling [12]. During skeletal muscle formation, the signal-responsive association with class IIa HDACs allows for a stagedependent control of MEF2 activity. Upon initiation of the differentiation program, the inhibitory activity of class IIa HDACs towards MEF2 is overcome by phosphorylation of their conserved serine residues by myogenic kinases [12,13]. This derepression allows MEF2 to cooperate with MyoD-related myogenic factors and promote skeletal muscle differentiation in vitro [11,14]. Historically, the role of the class IIa HDACs/MEF2 axis has thus been restricted to the early myoblastic differentiation events. Because the various approaches that were used to modulate the levels, activities or signal-responsiveness of class IIa HDACs had dramatic effects on myoblast differentiation [11-13,15], it has been very difficult so far to investigate any potential role of the class IIa/MEF2 pathway in later stages of skeletal muscle formation, such as myocyte fusion into myotubes.
The idea that the role of MEF2 during skeletal myogenesis was strictly limited to early differentiation is challenged by a careful survey of the literature. For instance, several studies have reported that MEF2 controls the transcription of genes involved in cytoskeleton structure and dynamics in muscle and non-muscle cells [16-20]. Cytoskeleton dynamics is required for proper cell-cell fusion, and accordingly many proteins important for myogenic fusion are components or regulators of the cytoskeleton (For recent review [21]). Based on these observations, one might expect that MEF2 may also control all or part of the fusion process during myogenesis.
MEF2 also Controls Myocyte Fusion
Few studies so far have provided direct evidence that MEF2 is a regulator of muscle cell fusion. In Drosophila, Arredondo et al. showed that CF2 (Drosophila Chorion factor 2), a MEF2 co-factor, is involved in the control of muscle size and of the number of nuclei in myofibers [22]. MEF2a-deficient murine C2C12 myoblasts have impaired ability to differentiate into myocytes but also exhibit lower levels of Myoferlin, which is critical for myogenic fusion, and of Palladin (Palld) and Formin-binding protein (Fnbp1l), which are involved in cytoskeleton dynamics and myocyte fusion [5]. Expression of singles bar gene in Drosophila, a protein linked to myocyte fusion, is also controlled by MEF2 [23].
The original goal of our study was to investigate the role of protein phosphatases during myogenesis [4]. Because protein kinases have long been known to be crucial in this process, the involvement of their counterpart, protein phosphatases would also be expected. Surprisingly, very few reports had addressed this issue when we started our project. Only three protein phosphatases had been shown to play a role in skeletal myogenesis and all three (i.e., calcineurin, protein phosphatase-1 and proteintyrosine phosphatase-α) were associated to myoblastic differentiation [24-26]. We decided to focus on PP2A because of the importance of class IIa HDACs during myogenesis and because of the PP2A ability to regulate these enzymes in other biological processes such as angiogenesis [27,28]. Our work showed that in myoblasts, PP2A controls HDAC4 phosphorylation levels and its repressive activity towards MEF2-directed transcription. PP2A is a trimeric holoenzyme made of a scaffolding A, catalytic C and B-type regulatory subunits. There are 15 different B-type subunits, that are classified into four distinct families (PR55/B or B55, PR61B’ or B56, PR72B” and STRN/PR93/PR110/B”’) [29,30].
The nature of the B subunit found in the holoenzyme determines various characteristics of the complex, including substrate specificity. Consequently, among the approximately 80 unique heterotrimeric holoenzymes, each possesses its own substrate specificity related to a particular biological activity. In muscle cells, we succeeded in identifying HDAC4 as the specific substrate of PP2A holoenzymes containing the Bδ regulatory subunit. Interestingly, other class IIa HDACs were not regulated by PP2A-Bδ [4].
To our surprise, knocking down the Bδ subunit had no significant effects on myoblast differentiation. Instead it severely impacted the ability of differentiated myocytes to fuse together and form multinucleated myofibers [4]. Using a transcriptomic approach, we confirmed that knockdown of PP2A-Bδ led to the up-regulation of MEF2-dependent genes. However, only a subset of MEF2- dependent genes, mainly those related to the cytoskeleton (i.e., Sorbs2) were affected by knocking down PP2A-Bδ, while those involved in myoblast differentiation (i.e. Myogenin) remained unaffected.
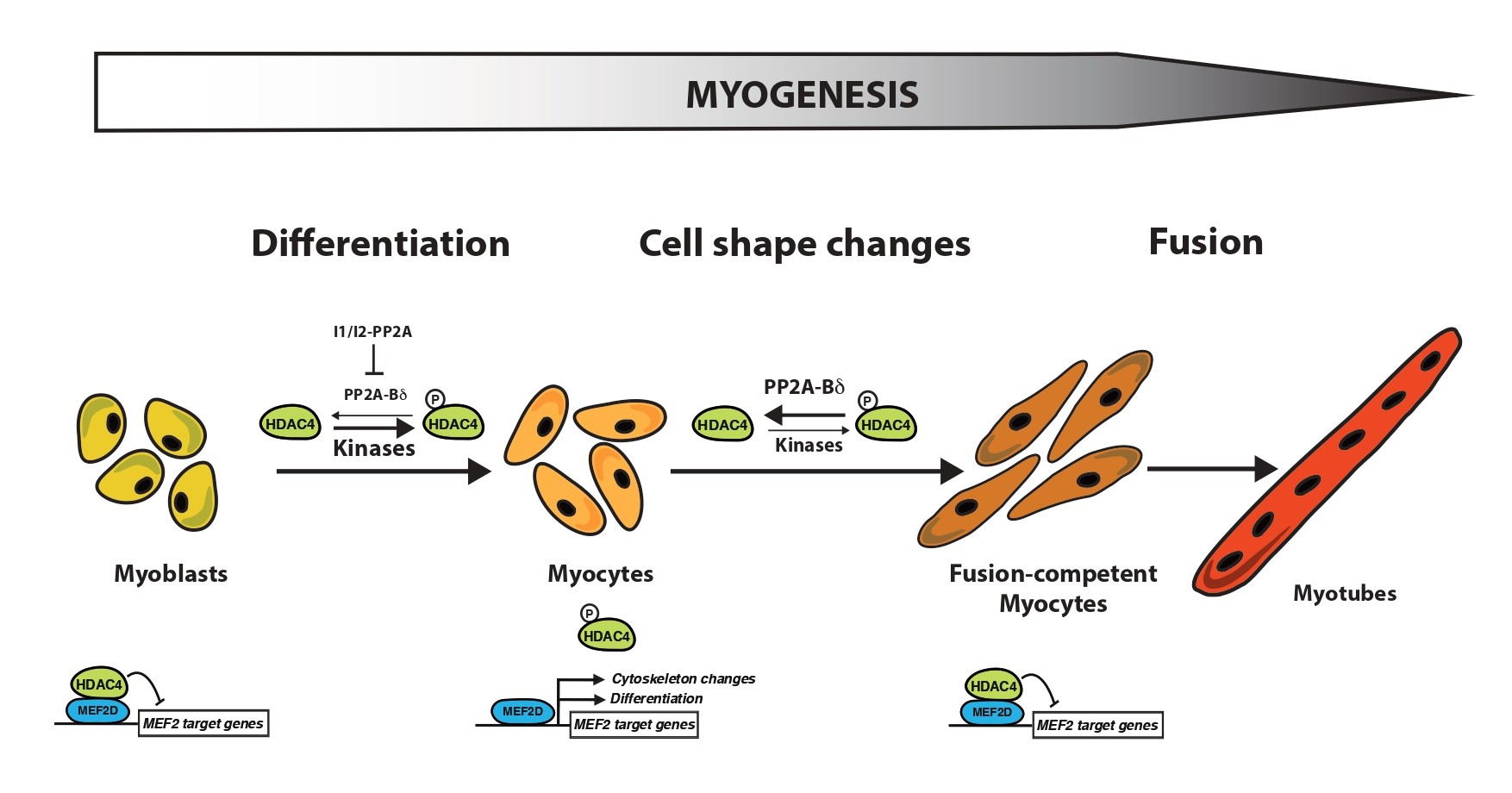
Some of our observations might provide a possible explanation as to why the absence of PP2A-Bδ has no impact on myocyte differentiation. Indeed, we found that I1PP2A and I2PP2A, two protein PP2A inhibitors [31], were highly expressed in non- and early differentiating myoblasts. It is likely that the presence of these proteins would prevent any PP2A activity during the early stages of myogenesis (Figure 1). The absence of phosphatase activity directed towards class IIa HDACs would create a favorable context for phosphorylation of HDAC4 by myogenic kinases, which would in turn activate the MEF2-directed myoblast differentiation program. At the onset of myogenic fusion, levels of I1PP2A and I2PP2A are drastically reduced, thus allowing a fine control of HDAC4 phosphorylation levels by PP2A-Bδ-mediated dephosphorylation.
A few months after our study was published, an article reported that HDAC4 expression levels were inversely correlated with the ability of myocytes to fuse and form myotubes [32]. In 2010, Sun et al. found that a mTOR–miR- 1–HDAC4–follistatin pathway was important for myocyte fusion [33]. In addition, mTOR and HDAC4 have both been connected to the endolysosomal pathway, which itself might be linked to myocyte fusion [34,35]. Knockdown of lysosomal associated protein 2 (LAMP-2), a key regulator of endolysomal biology, impairs myotube formation in C2C12 cells and LAMP-2 deficiency is associated with myopathies [35,36]. HDAC4 may regulate endolysosomal pH, a key feature in endolysosomal function, through the CREB-NEH6 axis [35]. Together with our observations, these studies point towards a key role for HDAC4 in the process of myogenic fusion. Autophagy, a lysosomemediated degradation pathway is activated during muscle cell differentiation and is required for in vitro myogenesis [36,37]. With regards to fusion, studies have shown that impaired autophagy is associated with reduced myotube formation [37], while others have reported opposite observations . Intriguingly, PP2A, HDAC4 and MEF2 have all been independently linked to the autophagic response [17,35,38,39]. It is thus tempting to speculate that in addition to cytoskeleton regulation, the PP2A-HDAC4- MEF2 axis may also regulate myocyte fusion through activation of alternative pathways, including autophagy. However, we did not observe upregulation of autophagyrelated genes in PP2A-Bδ-deficient myocytes.
Collectively, our data refine the historical role of the HDAC4-MEF2 pathway in skeletal myogenesis and shed light on its ability to control both the differentiation and the fusion processes. Nevertheless, our work also leaves a number of unanswered questions: Are there any phosphatase(s) controlling the phosphorylation of other class IIa HDACs that are also expressed in muscle cells (i.e. HDAC5, HDAC7 and HDAC9) [40]. Which processes, during myogenesis, are controlled by these unidentified phosphatases: differentiation, fusion, or both? What are the sets of MEF2-dependent genes that are specifically regulated by HDAC5, HDAC7 or HDAC9? Along this line, an exciting possibility would be that each class IIa HDAC phosphatase might ensure the control of specific subsets of MEF2-dependent genes. Beyond myogenesis, the question as to whether class IIa HDACs or MEF2 might also contribute to cell fusion in other biological processes requires further examination.
Interestingly, besides myogenesis, the control of cytoskeleton dynamics by the PP2A-MEF2 axis has also been linked to Alzheimer’s disease (AD). Tau, a microtubule-binding protein is hyperphosphorylated in amyloid plaques. This is a hallmark of AD and interestingly, PP2A-Bα is the major phosphatase regulating Tau phosphorylation [41]. Misregulation of PP2A activity leads to Tau hyperphosphorylation and aberrant microtubule dynamics. Therefore, pharmacological modulation of PP2A activity in AD has attracted a lot of attention [42]. To our knowledge, the role of PP2A-Bδ remains unexplored in neurodegenerative diseases, including AD. In contrast, mutations in MEF2c have also been associated to lateonset of AD while recent observations pointed to a role for HDAC4 in amyloid beta clearance in ApoE4 astrocytes [43-45]. Based on our observations in myocytes, we believe that the potential involvement of the PP2A-Bδ-HDAC4- MEF2 pathway in AD is an interesting line of investigation.
Therapeutic Perspectives
Protein phosphorylation is a key regulatory mechanism that is involved in virtually every signaling pathway in mammalian cells. Therefore, it is not surprising that dysregulation of protein phosphorylation has been linked to a plethora of diseases, such as cancer, inflammatory syndromes and Alzheimer’s disease to name only a few [46]. As of 2018, 38 kinase inhibitors had been approved for therapeutic use [47]. Although phosphatases are the opposing arm of protein phosphorylation regulation, they caught less attention than kinases as drug targets and the identification of phosphatase physiological substrates often remains a challenge [48]. For a long time, phosphatases were considered as non-optimal for pharmacological targeting because they were wrongly regarded as enzymes lacking specificity. Even if the growing knowledge about phosphatases and their substrates is slowing shifting this outdated paradigm, the development of phosphatasetargeting drugs to modulate phosphorylation levels of disease proteins remains mostly uncharted territory.
Targeting PP2A is even more challenging because of the involvement of PP2A in a wide range of key biological processes. Potential drugs thus need to target only specific PP2A holoenzymes, in order to avoid major adverse effects, as illustrated by the lethal knock-down of PP2A catalytic subunits in mice [49]. Recently, Leonard et. al identified DT-061, a compound that specifically stabilizes the B56/ B’α-PP2A holoenzyme in an active state [50]. This finding paves the way towards the development of innovative therapeutics targeting PP2A holoenzymes [51]. In light of our findings, a drug that would lead to the specific activation of PP2A-Bδ holoenzyme may be useful to favor muscle regeneration in the multiple diseases associated with muscle loss, i.e. sarcopenia or myopathies.
Conclusions and Future Directions
Although MEF2 is undisputedly important for myogenesis in vitro, its role in in vivo models is far less characterized and requires further investigation [3]. Indeed, MEF2 seems to be more prominent for myogenesis in Drosophila than in vertebrates. In mouse and Zebrafish, MEF2 is important for muscle regeneration and sarcomere assembly but seems dispensable for myotome formation. The discrepancies between in vivo and in vitro observations may lie in the functional redundancies between MEF2 members or splicing isoforms.
Interestingly, the phenotype that we observed after PP2A-Bδ knockdown was similar both in vitro (C2C12 mouse myoblasts model) and in vivo (zebrafish model). This suggests that interfering with regulators or partners might be a more profitable strategy when attempting to specifically modulate MEF2 biological functions.
Competing and Declaration of Interests
All listed authors have approved the manuscript and agreed with the submission. In addition, we declare no financial interest related to this work.
Funding
T.C was supported by grants from the Agence Nationale de la Recherche (Labex LipSTIC, ANR-11- LABX-0021) and the Région de Bourgogne Franche-Comté (support to Labex LipSTIC [to PS] 2017). TC received financial support from the Région de Bourgogne Franche-Comté (postdoctoral fellowship).
F.D was supported by the Interuniversity Attraction Poles Program initiated by the Belgian Science Policy Office (IUAP-BELSPO PVI/28 and PVII/13) and was supported by the Belgian National Fund for Scientific Research, the Belgian Foundation against Cancer (FAF-F/2018/1312) and Funds from the Université de Liège and the Fondation Léon Fredericq.
References
2. Hindi SM, Tajrishi MM, Kumar A. Signaling mechanisms in mammalian myoblast fusion. Sci Signal. 2013 Apr 23;6(272):re2.
3. Taylor MV, Hughes SM. Mef2 and the skeletal muscle differentiation program. Semin Cell Dev Biol. 2017 Nov 22;72:33-44.
4. Veloso A, Martin M, Bruyr J, O’Grady T, Deroanne C, Mottet D, et al. Dephosphorylation of HDAC4 by PP2A-Bd unravels a new role for the HDAC4/MEF2 axis in myoblast fusion. Cell Death Dis. 2019 Jul 4;10(7):512.
5. Estrella NL, Desjardins CA, Nocco SE, Clark AL, Maksimenko Y, Naya FJ. MEF2 transcription factors regulate distinct gene programs in mammalian skeletal muscle differentiation. J Biol Chem. 2015 Jan 9;290(2):1256-1268.
6. Asfaha Y, Schrenk C, Alves Avelar LA, Hamacher A, Pflieger M, Kassack MU, et al. Recent advances in class IIa histone deacetylases research. Bioorg Med Chem. 2019 Sep 6;115087.
7. Potthoff MJ, Olson EN. MEF2: a central regulator of diverse developmental programs. Development. 2007 Dec;134(23):4131-4140.
8. Haberland M, Montgomery RL, Olson EN. The many roles of histone deacetylases in development and physiology: implications for disease and therapy. Nat Rev Genet. 2009 Jan;10(1):32-42.
9. Mathias RA, Guise AJ, Cristea IM. Post-translational modifications regulate class IIa histone deacetylase (HDAC) function in health and disease. Mol Cell Proteomics. 2015 Mar;14(3):456–470.
10. Di Giorgio E, Brancolini C. Regulation of class IIa HDAC activities: it is not only matter of subcellular localization. Epigenomics. 2016 Feb;8(2):251-269.
11. Lu J, McKinsey TA, Zhang CL, Olson EN. Regulation of skeletal myogenesis by association of the MEF2 transcription factor with class II histone deacetylases. Mol Cell. 2000 Aug;6(2):233-244.
12. Simmons BJ, Cohen TJ, Bedlack R, Yao T-P. HDACs in skeletal muscle remodeling and neuromuscular disease. Handb Exp Pharmacol. 2011;206:79-101.
13. Knight JD, Kothary R. The myogenic kinome: protein kinases critical to mammalian skeletal myogenesis. Skelet Muscle. 2011 Sep 8;1:29.
14. Black BL, Molkentin JD, Olson EN. Multiple roles for the MyoD basic region in transmission of transcriptional activation signals and interaction with MEF2. Mol Cell Biol. 1998 Jan;18(1):69-77.
15. McKinsey TA, Zhang CL, Olson EN. Activation of the myocyte enhancer factor-2 transcription factor by calcium/calmodulin-dependent protein kinase-stimulated binding of 14-3-3 to histone deacetylase 5. Proc Natl Acad Sci USA. 2000 Dec 19;97(26):14400-14405.
16. Estrella NL, Naya FJ. Transcriptional networks regulating the costamere, sarcomere, and other cytoskeletal structures in striated muscle. Cell Mol Life Sci. 2013 Nov 12;71(9):1641-1656.
17. Tobin SW, Yang D, Girgis J, Farahzad A, Blais A, McDermott JC. Regulation of Hspb7 by MEF2 and AP-1: implications for Hspb7 in muscle atrophy. J Cell Sci. 2016 Nov 1;129(21):4076-4090.
18. Wales S, Hashemi S, Blais A, McDermott JC. Global MEF2 target gene analysis in cardiac and skeletal muscle reveals novel regulation of DUSP6 by p38MAPK-MEF2 signaling. Nucleic Acids Res. 2015 Feb 1;42(18):11349- 11362.
19. Lu YW, Lowery AM, Sun L-Y, Singer HA, Dai G, Adam AP, et al. Endothelial myocyte enhancer factor 2c inhibits migration of smooth muscle cells through fenestrations in the internal elastic lamina. Arterioscler Thromb Vasc Biol. 2017 May 4;37(7):1380-1390.
20. Vanpoucke G, Goossens S, De Craene B, Gilbert B, van Roy F, Berx G. GATA-4 and MEF2C transcription factors control the tissue-specific expression of the alphaT-catenin gene CTNNA3. Nucleic Acids Res. 2004 Jan 1;32(14):4155-4165.
21. Chen B, You W, Wang Y, Shan T. The regulatory role of Myomaker and Myomixer-Myomerger- Minion in muscle development and regeneration. Cell Mol Life Sci. 2019 Oct 23;
22. Arredondo JJ, Vivar J, Laine-Menéndez S, Martínez- Morentin L, Cervera M. CF2 transcription factor is involved in the regulation of Mef2 RNA levels, nuclei number and muscle fiber size. PLoS One. 2017 Jun 15;12(6):e0179194.
23. Brunetti TM, Fremin BJ, Cripps RM. Identification of singles bar as a direct transcriptional target of Drosophila Myocyte enhancer factor-2 and a regulator of adult myoblast fusion. Dev Biol. 2015 May 15;401(2):299-309.
24. Friday BB, Mitchell PO, Kegley KM, Pavlath GK. Calcineurin initiates skeletal muscle differentiation by activating MEF2 and MyoD. Differentiation. 2003 Apr;71(3):217-227.
25. Lu H, Shah P, Ennis D, Shinder G, Sap J, Le-Tien H, et al. The differentiation of skeletal muscle cells involves a protein-tyrosine phosphatase-alpha-mediated C-Src signaling pathway. J Biol Chem. 2002 Nov 29;277(48):46687-46695.
26. Kim SS, Kim JH, Lee S-H, Chung SS, Bang O-S, Park D, et al. Involvement of protein phosphatase- 1-mediated MARCKS translocation in myogenic differentiation of embryonic muscle cells. J Cell Sci. 2002 Jun 15;115(Pt 12):2465-2473.
27. Martin M, Geudens I, Bruyr J, Potente M, Bleuart A, Lebrun M, et al. PP2A regulatory subunit Ba controls endothelial contractility and vessel lumen integrity via regulation of HDAC7. EMBO J. 2013 Sep 11;32(18):2491- 2503.
28. Martin M, Potente M, Janssens V, Vertommen D, Twizere J-C, Rider MH, et al. Protein phosphatase 2A controls the activity of histone deacetylase 7 during T cell apoptosis and angiogenesis. Proc Natl Acad Sci USA. 2008 Mar 25;105(12):4727-4732.
29. Sents W, Ivanova E, Lambrecht C, Haesen D, Janssens V. The biogenesis of active protein phosphatase 2A holoenzymes: a tightly regulated process creating phosphatase specificity. FEBS J. 2013 Jan;280(2):644- 661.
30. Seshacharyulu P, Pandey P, Datta K, Batra SK. Phosphatase: PP2A structural importance, regulation and its aberrant expression in cancer. Cancer Lett. 2013 Jul 10;335(1):9-18.
31. Li M, Guo H, Damuni Z. Purification and characterization of two potent heat-stable protein inhibitors of protein phosphatase 2A from bovine kidney. Biochemistry. 1995 Feb 14;34(6):1988-1996.
32. Adler K, Molina PE, Simon L. Epigenomic mechanisms of alcohol-induced impaired differentiation of skeletal muscle stem cells; role of Class IIA histone deacetylases. Physiol Genomics. 2019 Aug 9;
33. Sun Y, Ge Y, Drnevich J, Zhao Y, Band M, Chen J. Mammalian target of rapamycin regulates miRNA-1 and follistatin in skeletal myogenesis. J Cell Biol. 2010 Jun 28;189(7):1157-1169.
34. Liu GY, Sabatini DM. mTOR at the nexus of nutrition, growth, ageing and disease. Nat Rev Mol Cell Biol. 2020 Jan 14;21(4):183-203.
35. Prasad H, Rao R. Histone deacetylase-mediated regulation of endolysosomal pH. J Biol Chem. 2018 May 4;293(18):6721-6735.
36. Sin J, Andres AM, Taylor DJR, Weston T, Hiraumi Y, Stotland A, et al. Mitophagy is required for mitochondrial biogenesis and myogenic differentiation of C2C12 myoblasts. Autophagy. 2016;12(2):369-380.
37. Fortini P, Ferretti C, Iorio E, Cagnin M, Garribba L, Pietraforte D, et al. The fine tuning of metabolism, autophagy and differentiation during in vitro myogenesis. Cell Death Dis. 2016 Mar 31;7(3):e2168.
38. Castets P, Rion N, Théodore M, Falcetta D, Lin S, Reischl M, et al. mTORC1 and PKB/Akt control the muscle response to denervation by regulating autophagy and HDAC4. Nat Commun. 2019 Jul 18;10(1):3187.
39. Wong P-M, Feng Y, Wang J, Shi R, Jiang X. Regulation of autophagy by coordinated action of mTORC1 and protein phosphatase 2A. Nat Commun. 2015 Aug 27;6:8048.
40. Martin M, Kettmann R, Dequiedt F. Class IIa histone deacetylases: conducting development and differentiation. Int J Dev Biol. 2009;53(2-3):291-301.
41. Braithwaite SP, Stock JB, Lombroso PJ, Nairn AC. Protein phosphatases and Alzheimer’s disease. Prog Mol Biol Transl Sci. 2012;106:343-379.
42. Taleski G, Sontag E. Protein phosphatase 2A and tau: an orchestrated “Pas de Deux”. FEBS Lett. 2018;592(7):1079-1095.
43. Rosenthal SL, Kamboh MI. Late-Onset Alzheimer’s Disease Genes and the Potentially Implicated Pathways. Curr Genet Med Rep. 2014 Mar 22;2:85-101.
44. Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet. 2013 Dec;45(12):1452- 1458.
45. Prasad H, Rao R. Amyloid clearance defect in ApoE4 astrocytes is reversed by epigenetic correction of endosomal pH. Proc Natl Acad Sci USA. 2018 Jul 10;115(28):E6640-E6649.
46. Cohen P. The role of protein phosphorylation in human health and disease. The Sir Hans Krebs Medal Lecture. Eur J Biochem. 2001 Oct;268(19):5001-5010.
47. Ferguson FM, Gray NS. Kinase inhibitors: the road ahead. Nat Rev Drug Discov. 2018 Mar 16;17(5):353-377.
48. Brautigan DL. Protein Ser/Thr phosphatases--the ugly ducklings of cell signalling. FEBS J. 2013 Jan;280(2):324- 345.
49. Reynhout S, Janssens V. Physiologic functions of PP2A: Lessons from genetically modified mice. Biochim Biophys Acta Mol Cell Res. 2019;1866(1):31-50.
50. Leonard D, Huang W, Izadmehr S, O’Connor CM, Wiredja DD, Wang Z, et al. Selective PP2A Enhancement through Biased Heterotrimer Stabilization. Cell. 2020 Apr 30;181(3):688-701.e16.
51. Sangodkar J, Farrington CC, McClinch K, Galsky MD, Kastrinsky DB, Narla G. All roads lead to PP2A: exploiting the therapeutic potential of this phosphatase. FEBS J. 2016 Mar;283(6):1004-1024.