Abstract
The Activating Molecule in Beclin-1-Regulated Autophagy (AMBRA1) is a scaffold protein involved in many cellular processes, including autophagy, apoptosis, cell growth and development. AMBRA1 functions as a substrate receptor of the DDB1-Cullin4-RBX1 ubiquitin E3 ligase complex that plays key roles in autophagy and the cell cycle regulatory network. Considering the crucial role of AMBRA1 in cellular homeostasis, structural and functional studies are important for understanding the mechanisms that coordinate these cell responses. Autophagy defects and impaired AMBRA1 function may contribute to the pathogenesis of several diseases, including cancer and neurodegenerative disorders. As a result, targeting AMBRA1 has gained interest as a potential therapeutic strategy. Due to the intrinsic disorder of AMBRA1, its structure has not been fully elucidated. A report by Liu et al., provided new insights into the structure and function of AMBRA1 [1]. This mini-review aims to summarize the DDB1-AMBRA1 complex structure and regulatory mechanism and discuss future research directions.
Keywords
Autophagy, Cell-cycle control, Ubiquitination, Tumorigenesis, AMBRA1, E3 ligase
Main Text
Two major intracellular protein degradation pathways, the ubiquitin–proteasome system (UPS) and autophagy are crucial for protein quality control in eukaryotic cells [2]. In general, short-lived and damaged proteins, such as denatured, unfolded or misfolded proteins are targeted by UPS, whereas long-lived proteins, insoluble protein aggregates and organelles are degraded by autophagy. The UPS system depends on ubiquitination by the addition of a small protein ubiquitin to specific lysine residues on target proteins. Ubiquitination is a sequential three-step reaction involving E1 (a ubiquitin-activating enzyme), E2 (a ubiquitin-conjugating enzyme) and E3 (a ubiquitin ligase enzyme), resulting in covalent attachment of ubiquitin to the target protein. The human genome contains two E1 genes and approximately 40 E2 genes. Additionally, there are over 600 E3 ligase genes, and a single E3 ligase may target multiple substrates [3-5]. E3 ligases can be classified into four families: the HECT, RING-finger, U-box and RBR families [6]. The RING-finger family is the most abundant E3 ligase family and is characterized by the presence of a zinc-binding domain called RING (Really Interesting New Gene) [7]. The Cullin4-DDB1-RBX1 complex is a classic member of the RING-finger family [8]. Cullin4 (CUL4) serves as a scaffold, its N-terminus binds to a RING-box protein (RBX1) which brings the E2 and the substrate into close proximity, while the C-terminus interacts with an adaptor protein DDB1 (damage-specific DNA binding protein 1). DDB1 can recruit substrates to the E3 ligase by interacting with different substrate receptors known as DCAFs (CUL4-associated factors).
The activating molecule in Beclin-1-regulated autophagy (AMBRA1) is a DCAF and is reported to bind DDB1 as a substrate receptor for the CRL4 E3 ligase [9]. AMBRA1 was identified as an autophagy-associated protein that coordinates different cellular process including cell proliferation, autophagy, UPS, apoptosis and mitophagy [10-14]. AMBRA1 contains an N-terminal WD40 domain which mediates interaction with DDB1. In addition, AMBRA1 sequence contains a large proportion of intrinsically unstructured regions (IDR). The conformational flexibility of AMBRA1 IDRs may facilitate multiple protein-protein interactions required for coordination of several biological processes [14,15]. AMBRA1 can positively regulate Beclin1-dependent autophagy through directly binding to Beclin1, which is needed for neurogenesis during the embryonic period [16,17]. Beclin1 and VPS34 are the components of the class III phosphatidylinositol-3-kinase complex, which initiates autophagosome formation by producing phosphatidylinositol-3-phosphate (PI3P) [18,19]. Mechanistically, AMBRA1 mediates K63-linked ubiquitylation of Beclin1, thereby promoting the interaction between Beclin1 and VPS34 and autophagy [20,21]. AMBRA1 is regulated by post-translational modifications, including phosphorylation and ubiquitination. Upon autophagy induction, Unc-51-like autophagy-activating kinase 1 (ULK1) phosphorylates AMBRA1 to promote it release from microtubules by inhibiting AMBRA1 interaction with dynein light chain (DLC)1 [22,23]. Subsequently, AMBRA1 re-localizes to the ER to enable autophagosome formation [23-25]. AMBRA1 also can serve as the substrate of CUL4-DDB1 and RNF2 (The ring finger protein 2) E3 ligase complexes which can mediate its ubiquitination and degradation [9,26].
Three recent reports have revealed that AMBRA1 mediates ubiquitination and degradation of cyclin D1 (Figure 1a) [13,27,28]. Cyclin D1 is a cell cycle regulator that controls G1 to S phase progression in many cell types, and a high expression level of this gene is related to cancer development [29]. Mechanistically, cyclin-dependent kinases CDK4/6 bind to cyclin D1 to form an active complex that phosphorylates and inactivates the tumor suppressor protein retinoblastoma protein (RB) [30-32]. Phosphorylated RB dissociates from the E2F transcription factor, and E2F regulates transcription of genes necessary for progression through the S phase and promotes cell cycle progression [33,34]. Loss of AMBRA1 leads to increased cellular levels of cyclin D1, resulting in increased cell proliferation, replication stress and tumorigenic activity [13,27,28] (Figure 1a). Moreover, loss of AMBRA1 promotes the formation of cyclin D1/CDK2 complexes, resulting in decreased sensitivity to CDK4/6 inhibitors [28]. These findings highlight the need for combined inhibition of CDK2/4/6 in cancer therapy. In addition to cyclin D1, AMBRA1 can regulate cell proliferation by activating protein phosphatase 2A (PP2A). PP2A dephosphorylates the proto-oncogene c-Myc at Ser62, thereby destabilizing c-Myc and decreasing cell proliferation and tumorigenesis (Figure 1b) [10,35]. Taken together, AMBRA1 functions as the platform for extensive crosstalk between cell cycle, proliferation, and autophagy through multiple protein-protein interactions.
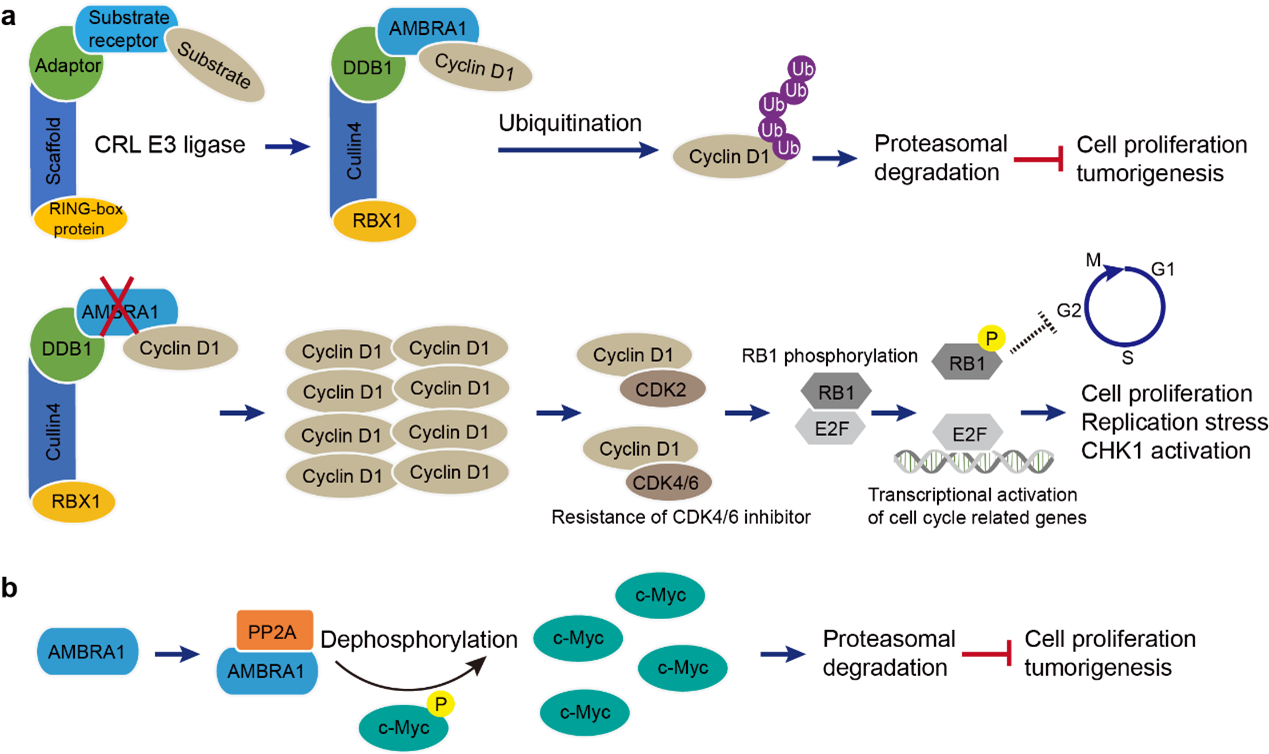
Figure 1. AMBRA1 links to cell cycle and tumorigenic activity. a. Cartoon schematic of the Cullin4-DDB1-RBX1-AMBRA1 complex and its regulation of the cell cycle and cell proliferation by ubiquitination of cyclin D1. The absence of AMBRA1 stabilizes cyclin D1, and results in the formation of the cyclin D1-CDK2 and cyclin D1-CDK4/6 complex, which phosphorylates RB1 and induces cell proliferation. b. The AMBRA1-PP2A complex activates the dephosphorylation of PP2A and destabilizes c-Myc.
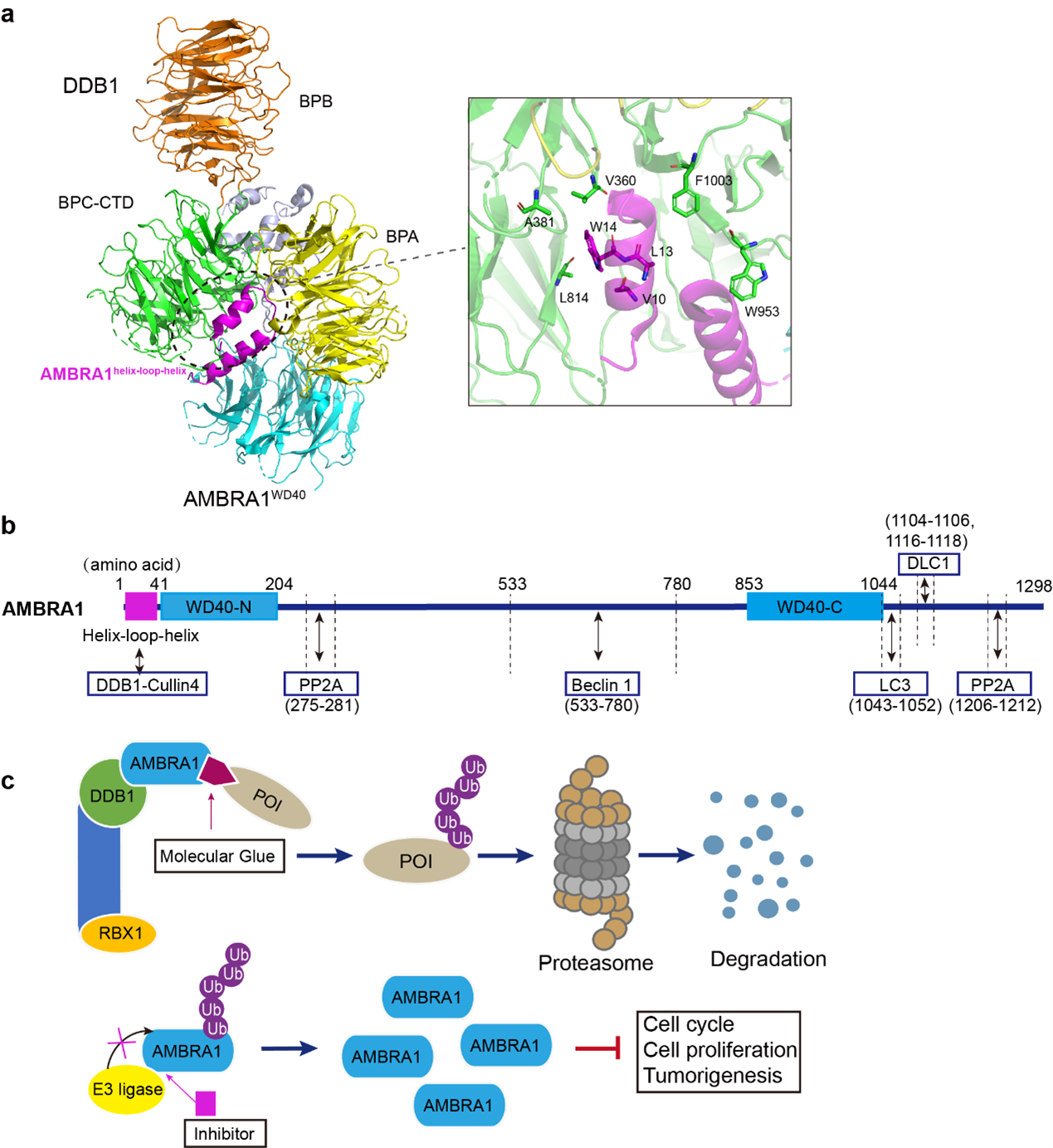
Understanding the structural basis of the DDB1-AMBRA1 WD40 complex is essential for elucidating its functional and regulatory mechanisms as well as exploring its therapeutic potential [36,37]. A recent study by Liu et al. determined the cryo-EM structure of the human DDB1-AMBRA1 WD40 complex and revealed a “split” WD40 domain formed by the N-terminal (residues 1-204) and C-terminal (residues 853–1044) regions [1]. DDB1 is composed of three seven-bladed WD40 β-propeller domains, BPA, BPB, BPC, and a C-terminal helical domain (CTD). The BPA and BPC domains are arranged in a double-β-propeller, forming a binding pocket for substrate receptors at the interface (Figure 2a). This DDB1 interface pocket is where the AMBRA1 WD40 domain binds through the N-terminal helix-loop-helix motif. The key residues of AMBRA1 participating in the interaction with DDB1 include V10, L13 and W14 (Figure 2a). The CUL4-RBX1-DDB1-AMBRA1 WD40 is able to bind the substrate Cyclin D1/CDK4 complex and mediate its ubiquitination. Mutating the key binding residues V10R and L13R in AMBRA1 disrupts the interaction with DDB1 and prevents ubiquitination of Cyclin D1. Disrupting the interaction between DDB1 and AMBRA1 or knocking down AMBRA1 dysregulates the cell cycle control and results in genomic instability [1,13,27,28]. Finally, interaction between AMBRA1 and CRL4 E3 ligase complex is essential for the induction of autophagy and mutations that disrupt the DDB1-AMBRA1 WD40 interaction interface impair autophagy [38]. One notable feature of AMBRA1 is the presence of intrinsically disordered regions within its structure. These regions lack well-defined three-dimensional structures and exhibit high flexibility under the native physiologic conditions. This flexibility allows AMBRA1 to undergo conformational changes and function as a receptor protein in various cellular complexes. The stabilization of the AMBRA1 WD40 domain by DDB1 likely serves to organize and bring together various substrates and CRL4 E3 ligase. This process assists in facilitating the transfer of ubiquitin, enabling AMBRA1 to coordinate multiple biological processes in response to different inputs. Recent studies of the AMBRA1 WD40-CRL4 E3 ligase complex provide a structural context in which to test how AMBRA1 loss can lead to uncontrolled cell growth and promote tumorigeneses and paving the way for development of novel therapeutic strategies targeting autophagy and cell cycle dysregulation in human diseases.
Conclusion and Future Perspectives
AMBRA1 functions as a scaffold protein to coordinate several cellular processes, including the cell cycle, cell proliferation, neurogenesis, autophagy and the UPS, and has wide implications for a variety of diseases in humans. AMBRA1 acts as a tumor suppressor and regulates two parallel pathways, the c-Myc pathway and the cyclin D–CDK4/6–RB–E2F pathway, which drive the G1–S transition [39]. Decreased AMBRA1 expression levels lead to accumulation of cyclin D, promote cell division and result in desensitization of CDK4/6 inhibitors [13,27,28]. RNF2 E3 ligase in complex with WASH can mediate the ubiquitination and degradation of AMBRA1 [9,26].
It is reported that AMBRA1 plays a key role in tumorigenesis and progression [40]. For example, deficiency of AMBRA1 promotes melanoma growth and invasion [11], induces lung adenocarcinoma in a mouse model, and is correlated with overall worse survival in lung adenocarcinoma patients [13]. Therefore, defects in AMBRA1 might be useful as cancer progression biomarker in oncology. In addition to the identified interactors, such as Beclin1 [41], LC3 [42], cyclin D1[13] and PP2A [10] (Figure 2b), a comprehensive proteomic study is needed to unravel the intricate network of AMBRA1 interactions and post-translational modifications. Molecular details of AMBRA1 interactions with different substrates and regulatory proteins are required to fully understand the substrate specificity and for the design of small molecule drugs targeting protein-protein interactions. Furthermore, it would be important to gain a deeper understanding of how cells regulate AMBRA1 expression levels and intracellular distribution and explore potential therapeutic strategies.
The recently reported structure of the DDB1-AMBRA1 WD40 E3 ligase receptor complex provides the basis for further investigations into its therapeutic potential [1]. The molecular details of protein-protein interaction interfaces provide valuable information for identification of new substrates that bind to DDB1-AMBRA1 WD40 and designing of small-molecule drugs. Targeted protein degradation (TPD) is an emerging therapeutic strategy to target dysregulated proteins that play key roles in diseases such as cancer and neurodegeneration. Molecular glues are small molecules that stabilize the interactions between proteins and can be used to reduce cellular levels of disease-causing proteins through TPD [43], enhance the interaction between protein of interest (POI) and the E3 ligase [44,45] and induce POI ubiquitination and degradation [46]. In addition, inhibitors could be designed to disrupt the interaction between E3 ligases which ubiquitinate AMBRA1, to decrease its degradation and potentially stimulate autophagy and/or maintain the cell cycle and proliferation at normal levels (Figure 2c). Whether some of these therapeutic strategies could be applied to AMBRA1 requires further research.
Figure 2. Model of the DDB1-AMBRA1WD40 complex and potential small molecule compound development. a. Model of the DDB1-AMBRA1WD40 complex and the binding interface (PDB 8WQR) [1]. The DDB1 domain and AMBRA1WD40 are colored as follows: BPA domain, yellow; BPB domain, orange; BPC-CTD domain, green and light blue; AMBRA1 WD40, cyan. b. The identified proteins that interact with AMBRA1 are shown in the corresponding binding region. c. The development of potential small-molecule compounds, including molecular glues and inhibitors. Molecular glue may enhance the interaction between the E3 ligase complex and the protein of interest (POI) and promote its degradation; the inhibitor interrupts the interaction between AMBRA1 and E3 ligase, reducing the ubiquitination and degradation of AMBRA1.
AMBRA1 plays a central role in the pathogenesis of multiple system atrophy (MSA) through interaction with α-synuclein and mediate its degradative dynamics. This raises the possibility that molecular modulation targeting AMBRA1 can be a promising candidate for the treatment of synucleinopathies [47]. Small molecules could be as well used to activate AMBRA1 by disrupting protein-protein interactions, such as DLC1 [22,23]. Disrupting the interaction between AMBRA1 and DLC1 can protect cells against apoptosis, induce autophagy and mitophagy in SHSY5Y cells, and enhance the switch from quiescence to proliferation in mouse neural stem cells. This strategy provides the potential to treat neurodegeneration [22]. Artificial Intelligence (AI) and Machine Learning (ML) algorithms have great potential for developing new drugs by improving efficiency, accuracy and speed of drug screening and discovery [48-50]. The employment of AI-driven techniques such as AlphaFold has greatly aided in prediction and structure determination of AMBRA1, thereby offering deeper insights into its functional mechanisms and opening up new avenues for innovative therapeutic strategies [1,51,52]. The advancement of deep learning algorithms has revolutionized the study of proteins, empowering precise predictions of intricate structures of proteins, nucleic acids, and small molecules [53]. These computational approaches hold great promise in the field of drug discovery and development. AMBRA1 research is still evolving, and the development of therapeutic interventions targeting AMBRA1 is in the early stages. Further studies are necessary to fully understand the potential of AMBRA1 as a therapeutic target and to evaluate its safety and efficacy in various disease contexts.
|
Interactors |
Binding region on AMBRA1 (based on canonical UniProt sequence) |
Reference |
|
Beclin1 |
Residue 533-780 |
[16,54] |
|
DDB1-Cullin4 complex |
Residue 1-41 |
[1] |
|
PP2A |
Residue 275-281 and 1206-1212 |
[10] |
|
DLC1 |
Residue 1104-1106 and 1116-1118 |
[23] |
|
TRAF6 |
Residue 618-623 and 681-686 |
[55] |
|
Elongin C-Cullin5 complex |
Residue 825-1298 |
[56] |
|
LC3 |
Residue 1043-1052 |
[42] |
|
ERLIN1 |
Residue 533-780 and 796-1298 |
[57] |
|
ULK1 |
Residue 1-532 and 781-1298 |
[55,58] |
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31950410540 to G.S.), the Foreign Young Talent Program from State Administration of Foreign Experts Affairs (QN2021032004L to G.S.), and the CUHK-Shenzhen University Development Fund (to G.S.).
Conflict of Interest
The authors declare that they have no conflicts of interest.
References
2. Kwon YT, Ciechanover A. The ubiquitin code in the ubiquitin-proteasome system and autophagy. Trends in Biochemical Sciences. 2017 Nov 1;42(11):873-86.
3. Pickart CM, Eddins MJ. Ubiquitin: structures, functions, mechanisms. Biochimica et Biophysica Acta (BBA)-Molecular Cell Research. 2004 Nov 29;1695(1-3):55-72.
4. Rotin D, Kumar S. Physiological functions of the HECT family of ubiquitin ligases. Nature Reviews Molecular Cell Biology. 2009 Jun;10(6):398-409.
5. Stewart MD, Ritterhoff T, Klevit RE, Brzovic PS. E2 enzymes: more than just middle men. Cell Research. 2016 Apr;26(4):423-40.
6. Yang Q, Zhao J, Chen D, Wang Y. E3 ubiquitin ligases: styles, structures and functions. Molecular Biomedicine. 2021 Dec;2:23.
7. Freemont PS, Hanson IM, Trowsdale J. A novel gysteine-rich sequence motif. Cell. 1991;64(3):483-4.
8. Lee J, Zhou P. DCAFs, the missing link of the CUL4-DDB1 ubiquitin ligase. Molecular Cell. 2007 Jun 22;26(6):775-80.
9. Antonioli M, Albiero F, Nazio F, Vescovo T, Perdomo AB, Corazzari M, et al. AMBRA1 interplay with cullin E3 ubiquitin ligases regulates autophagy dynamics. Developmental Cell. 2014 Dec 22;31(6):734-46.
10. Cianfanelli V, Fuoco C, Lorente M, Salazar M, Quondamatteo F, Gherardini PF, et al. AMBRA1 links autophagy to cell proliferation and tumorigenesis by promoting c-Myc dephosphorylation and degradation. Nature Cell Biology. 2015 Jan;17(1):20-30.
11. Di Leo L, Bodemeyer V, Bosisio FM, Claps G, Carretta M, Rizza S, et al. Loss of Ambra1 promotes melanoma growth and invasion. Nature Communications. 2021 May 5;12(1):2550.
12. Di Rita A, Peschiaroli A, D′ Acunzo P, Strobbe D, Hu Z, Gruber J, et al. HUWE1 E3 ligase promotes PINK1/PARKIN-independent mitophagy by regulating AMBRA1 activation via IKKα. Nature Communications. 2018 Sep 14;9(1):3755.
13. Chaikovsky AC, Li C, Jeng EE, Loebell S, Lee MC, Murray CW, et al. The AMBRA1 E3 ligase adaptor regulates the stability of cyclin D. Nature. 2021 Apr 29;592(7856):794-8.
14. Cianfanelli V, De Zio D, Di Bartolomeo S, Nazio F, Strappazzon F, Cecconi F. Ambra1 at a glance. Journal of Cell Science. 2015 Jun 1;128(11):2003-8.
15. Babu MM. The contribution of intrinsically disordered regions to protein function, cellular complexity, and human disease. Biochemical Society Transactions. 2016 Oct 15;44(5):1185-200.
16. Maria Fimia G, Stoykova A, Romagnoli A, Giunta L, Di Bartolomeo S, Nardacci R, et al. Ambra1 regulates autophagy and development of the nervous system. Nature. 2007 Jun 28;447(7148):1121-5.
17. Cecconi F, Piacentini M, Fimia GM. The involvement of cell death and survival in neural tube defects: a distinct role for apoptosis and autophagy?. Cell Death & Differentiation. 2008 Jul;15(7):1170-7.
18. Suzuki K, Ohsumi Y. Molecular machinery of autophagosome formation in yeast, Saccharomyces cerevisiae. FEBS Letters. 2007 May 22;581(11):2156-61.
19. Itakura E, Kishi C, Inoue K, Mizushima N. Beclin 1 forms two distinct phosphatidylinositol 3-kinase complexes with mammalian Atg14 and UVRAG. Molecular Biology of the Cell. 2008 Dec;19(12):5360-72.
20. Xia P, Wang S, Du Y, Zhao Z, Shi L, Sun L, et al. WASH inhibits autophagy through suppression of Beclin 1 ubiquitination. The EMBO Journal. 2013 Oct 16;32(20):2685-96.
21. Sun WL. Ambra1 in autophagy and apoptosis: Implications for cell survival and chemotherapy resistance. Oncology Letters. 2016 Jul 1;12(1):367-74.
22. Hawkins K, Watt M, Gillotin S, Hanspal M, Helley M, Richardson J, et al. Disrupting the interaction between AMBRA1 and DLC1 is a promising therapeutic strategy for neurodegeneration that prevents apoptosis while enhancing autophagy and mitophagy. BioRxiv. 2023:2023-10.
23. Di Bartolomeo S, Corazzari M, Nazio F, Oliverio S, Lisi G, Antonioli M, et al. The dynamic interaction of AMBRA1 with the dynein motor complex regulates mammalian autophagy. Journal of Cell Biology. 2010 Oct 4;191(1):155-68.
24. Fimia GM, Di Bartolomeo S, Piacentini M, Cecconi F. Unleashing the Ambra1-Beclin 1 complex from dynein chains: Ulk1 sets Ambra1 free to induce autophagy. Autophagy. 2011 Jan 1;7(1):115-7.
25. Tang DY, Ellis RA, Lovat PE. Prognostic impact of autophagy biomarkers for cutaneous melanoma. Frontiers in Oncology. 2016 Nov 9;6:236.
26. Xia P, Wang S, Huang G, Du Y, Zhu P, Li M, et al. RNF2 is recruited by WASH to ubiquitinate AMBRA1 leading to downregulation of autophagy. Cell Research. 2014 Aug;24(8):943-58.
27. Maiani E, Milletti G, Nazio F, Holdgaard SG, Bartkova J, Rizza S, et al. AMBRA1 regulates cyclin D to guard S-phase entry and genomic integrity. Nature. 2021 Apr 29;592(7856):799-803.
28. Simoneschi D, Rona G, Zhou N, Jeong YT, Jiang S, Milletti G, et al. CRL4AMBRA1 is a master regulator of D-type cyclins. Nature. 2021 Apr 29;592(7856):789-93.
29. Alao JP. The regulation of cyclin D1 degradation: roles in cancer development and the potential for therapeutic invention. Molecular Cancer. 2007 Dec;6:24.
30. Kato JY, Matsushime H, Hiebert SW, Ewen ME, Sherr CJ. Direct binding of cyclin D to the retinoblastoma gene product (pRb) and pRb phosphorylation by the cyclin D-dependent kinase CDK4. Genes & Development. 1993 Mar 1;7(3):331-42.
31. Lundberg AS, Weinberg RA. Functional inactivation of the retinoblastoma protein requires sequential modification by at least two distinct cyclin-cdk complexes. Molecular and Cellular Biology. 1998 Feb 1;18(2):753-761.
32. Weinberg RA. The retinoblastoma protein and cell cycle control. cell. 1995 May 5;81(3):323-30.
33. Bateman E. Autoregulation of eukaryotic transcription factors. Progress in nucleic acid Research and Molecular Biology. 1998 Jan 1;60:133-68.
34. Slansky JE, Farnham PJ. Introduction to the E2F family: protein structure and gene regulation. Transcriptional Control of Cell Growth: The E2F Gene Family. 1996 Jan 1:1-30.
35. Farrington CC, Yuan E, Mazhar S, Izadmehr S, Hurst L, Allen-Petersen BL, et.al. Protein phosphatase 2A activation as a therapeutic strategy for managing MYC-driven cancers. Journal of Biological Chemistry. 2020 Jan 17;295(3):757-770.
36. Blundell TL. Structure-based drug design. Nature. 1996 Nov 7;384(6604):23-6.
37. Özçelik R, van Tilborg D, Jiménez‐Luna J, Grisoni F. Structure‐Based Drug Discovery with Deep Learning. ChemBioChem. 2023 Jul 3;24(13):e202200776.
38. Liu M, Wang Y, Teng F, Mai X, Wang X, Su MY, et.al. Structural basis for substrate recruitment by AMBRA1 E3 ligase receptor. BioRxiv. 2022 Dec 4:2022-12.
39. Bartek J, Lukas J. Pathways governing G1/S transition and their response to DNA damage. FEBS Letters. 2001 Feb 16;490(3):117-22.
40. Li X, Lyu Y, Li J, Wang X. AMBRA1 and its role as a target for anticancer therapy. Frontiers in Oncology. 2022 Sep 27;12:946086.
41. Cianfanelli V, D'Orazio M, Cecconi F. AMBRA1 and BECLIN 1 interplay in the crosstalk between autophagy and cell proliferation. Cell Cycle. 2015 Apr 3;14(7):959-63.
42. Strappazzon F, Nazio F, Corrado M, Cianfanelli V, Romagnoli A, Fimia GM, et.al. AMBRA1 is able to induce mitophagy via LC3 binding, regardless of PARKIN and p62/SQSTM1. Cell Death & Differentiation. 2015 Mar;22(3):419-32.
43. Zhao L, Zhao J, Zhong K, Tong A, Jia D. Targeted protein degradation: mechanisms, strategies and application. Signal Transduction and Targeted Therapy. 2022 Apr 4;7(1):113.
44. Simonetta KR, Taygerly J, Boyle K, Basham SE, Padovani C, Lou Y, et.al. Prospective discovery of small molecule enhancers of an E3 ligase-substrate interaction. Nature Communications. 2019 Mar 29;10(1):1402.
45. Sasso JM, Tenchov R, Wang D, Johnson LS, Wang X, Zhou QA. Molecular glues: the adhesive connecting targeted protein degradation to the clinic. Biochemistry. 2022 Jul 20;62(3):601-23.
46. Békés M, Langley DR, Crews CM. PROTAC targeted protein degraders: the past is prologue. Nature Reviews Drug Discovery. 2022 Mar;21(3):181-200.
47. Miki Y, Tanji K, Mori F, Tatara Y, Utsumi J, Sasaki H, et.al. AMBRA1, a novel α‐synuclein‐binding protein, is implicated in the pathogenesis of multiple system atrophy. Brain Pathology. 2018 Jan;28(1):28-42.
48. Blanco-Gonzalez A, Cabezon A, Seco-Gonzalez A, Conde-Torres D, Antelo-Riveiro P, Pineiro A, et.al. The role of ai in drug discovery: challenges, opportunities, and strategies. Pharmaceuticals. 2023 Jun 18;16(6):891.
49. Sarkar C, Das B, Rawat VS, Wahlang JB, Nongpiur A, Tiewsoh I, et.al. Artificial intelligence and machine learning technology driven modern drug discovery and development. International Journal of Molecular Sciences. 2023 Jan 19;24(3):2026.
50. Paul D, Sanap G, Shenoy S, Kalyane D, Kalia K, Tekade RK. Artificial intelligence in drug discovery and development. Drug Discov Today. 2021 Jan;26(1):80-93.
51. Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et.al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021 Aug;596(7873):583-9.
52. Tiberti M, Di Leo L, Vistesen MV, Kuhre RS, Cecconi F, De Zio D, et.al. The Cancermuts software package for the prioritization of missense cancer variants: a case study of AMBRA1 in Melanoma. Cell Death & Disease. 2022 Oct 15;13(10):872.
53. Abramson J, Adler J, Dunger J, Evans R, Green T, Pritzel A, et.al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature. 2024 May 8:1-3.
54. Fimia GM, Corazzari M, Antonioli M, Piacentini M. Ambra1 at the crossroad between autophagy and cell death. Oncogene. 2013 Jul;32(28):3311-8.
55. Nazio F, Strappazzon F, Antonioli M, Bielli P, Cianfanelli V, Bordi M, et.al. mTOR inhibits autophagy by controlling ULK1 ubiquitylation, self-association and function through AMBRA1 and TRAF6. Nature Cell Biology. 2013 Apr;15(4):406-16.
56. Chen SH, Jang GM, Hüttenhain R, Gordon DE, Du D, Newton BW, et.al. CRL4AMBRA1 targets Elongin C for ubiquitination and degradation to modulate CRL5 signaling. The EMBO Journal. 2018 Sep 14;37(18):e97508.
57. Manganelli V, Matarrese P, Antonioli M, Gambardella L, Vescovo T, Gretzmeier C,et.al. Raft-like lipid microdomains drive autophagy initiation via AMBRA1-ERLIN1 molecular association within MAMs. Autophagy. 2021 Sep 2;17(9):2528-48.
58. Cianfanelli V, Nazio F, Cecconi F. Connecting autophagy: AMBRA1 and its network of regulation. Molecular & Cellular Oncology. 2015 Jan 2;2(1):e970059.