Keywords
Lipases, Cysteine-lipases, Catalytic triad, Enzyme activity, Digestive tract, Immunological response
Commentary
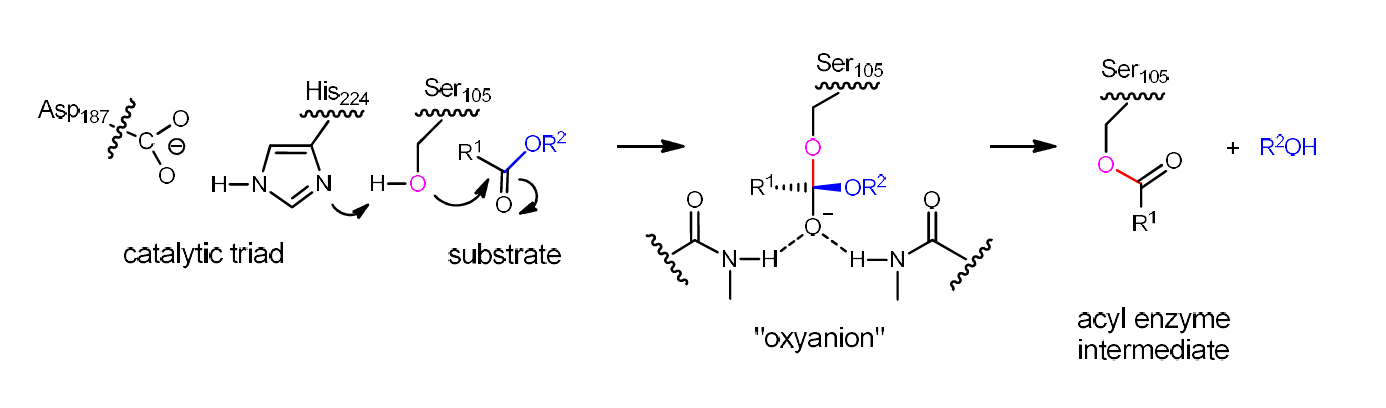
Lipases, esterases and proteases constitute superfamilies of hydrolases not only play an important role in the immune system, but also as catalysts in biotechnology and organic chemistry. Mechanistically, they all involve a similar catalytic triad. The mechanism of lipasecatalysis is defined by the catalytic triad Ser-His-Asp in which activated serine adds nucleophilically to the carbonyl function of an ester or lactone substrate in the rate-determining step with formation of a short-lived oxyanion which then fragments into an alcohol and a covalent acyl-enzyme intermediate, the latter rapidly undergoing reaction with water and liberating the respective carboxylic acid (Figure 1) [1-5].
In contrast, both serine-proteases with Ser-His-Asp as the catalytic triad and cysteine-proteases characterized by Cys-His-Asp have been identified as enzymes with high activity. It has also been shown that in the case of serine proteases, mutation to the respective cysteine-proteases leads to a partial or complete breakdown of activity, and the same applies to the opposite scenario in which a cysteine-protease is mutated into a serine-protease, which has led to lively discussions concerning the origin of these effects [6-12]. Recently, we demonstrated that the conversion of a serine-lipase into an artificial cysteinelipase also induces significant loss of activity [13].
Today, it is well known that the human digestive tract has a prominent influence on the immune system. As already alluded to in the above short introductory information, lipases play an important role in a number of health problems. Only a few typical studies are cited here, the focus in these cases being on such enzymes as monoacylglycerol lipases, triglycerol lipases, and phospholipases [14-26]. In these and other works available in the extensive literature, sequence information was generally presented, showing that serine-lipases are indeed involved; in some studies, this was just assumed by the authors and no mention of possible cysteinelipases as alternatives was made.
In our study describing the transformation of a serineto a cysteine-lipase, Candida antarctica lipase B (CALB) was used as the model hydrolase [13]. The catalytic triad of this standard and in biotechnology often applied lipase is Asp187-His-224-Ser105 (Figure 1) [1-5,27-28]. In our study, mutant Asp187-His-224-Cys105 as a cysteine-lipase was shown to have a very low activity for a number of structurally different substrates. In order to regain and perhaps even to surpass the activity of wildtype (WT) CALB in a model transformation involving the hydrolytic kinetic resolution of a racemic ester, we utilized the techniques of directed evolution [29-32]. Specifically, saturation mutagenesis at rationally chosen residues surrounding the binding pocket according to the Combinatorial Active-site Saturation Test (CAST) [33] was performed, followed by Iterative Saturation Mutagenesis (ISM) [33-34] at other hotspot residues around the binding pocket [13]. The best evolved mutant, W104V/S105C/A281Y/A282Y/V149G, showed a 40-fold enhancement of activity in the model reaction, and was even more active than WT CALB in the hydrolysis of further substrates.
The combination of X-ray structures, kinetics, molecular dynamics (MD) simulations and QM/MM computations revealed dynamic effects upon going from cysteine-CALB to the best mutant [13]. It was shown that the three additional mutations cause, inter alia, the re-adjustment of the now active catalytic triad Asp187-His-224-Cys105 in a way that produces the zwitterion Cys105-/His224+, thereby enforcing a novel 2-step mechanism rather than the traditional concerted addition process [13]. Changing the mechanism of an enzyme by introducing mutations is a rare event in protein engineering. The overall results demonstrate that cysteine-lipases can in fact be active, but they do not prove that such lipases occur in nature.
The distinction between lipases and esterases continues to be a subject of considerable interest. Traditionally it was believed that lipases have a lid which opens upon interaction with a hydrophobic substrate by interfacial activation, in contrast to esterases [1-5,35-36]. However, uncertainties remain, and a sharp distinction may not matter [37-38]. In view of our recent study [13], we believe that an intensive search for natural cysteine-lipases (or esterases) could be a fruitful venture in different areas of lipase research, immunology and virology.
References
2. Botos I, Wlodawer A. The expanding diversity of serine hydrolases. Current Opinion in Structural Biology. 2007 Dec 1;17(6):683-90.
3. Bornscheuer, UT, Kazlauskas, RJ. Hydrolases in Organic Synthesis. Wiley-VCH, 1999.
4. Schmid RD, Verger R. Lipases: interfacial enzymes with attractive applications. Angewandte Chemie International Edition. 1998 Jul 3;37(12):1608-33.
5. Melani NB, Tambourgi EB, Silveira E. Lipases: From production to applications. Separation & Purification Reviews. 2020 Apr 2;49(2):143-58.
6. Carter P, Wells JA. Dissecting the catalytic triad of a serine protease. Nature. 1988 Apr 7;332(6164):564-8.
7. Neet KE, Koshland Jr DE. The conversion of serine at the active site of subtilisin to cysteine: a” chemical mutation”. Proceedings of the National Academy of Sciences of the United States of America. 1966 Nov;56(5):1606.
8. Polgar L, Bender ML. A new enzyme containing a synthetically formed active site. Thiol-Subtilisin1. Journal of the American Chemical Society. 1966 Jul;88(13):3153- 4.
9. Higaki JN, Evnin LB, Craik CS. Introduction of a cysteine protease active site into trypsin. Biochemistry. 1989 Nov 1;28(24):9256-63.
10. Slade A, Horrocks AJ, Lindsay CD, Dunbar B, Virden R. Site-directed chemical conversion of serine to cysteine in penicillin acylase from Escherichia coli ATCC 11105: Effect on conformation and catalytic activity. European Journal of Biochemistry. 1991 Apr;197(1):75-80.
11. Wilke ME, Higaki JN, Craik CS, Fletterick RJ. Crystal structure of rat trypsin-S195C at− 150 C: Analysis of low activity of recombinant and semisynthetic thiol proteases. Journal of Molecular Biology. 1991 Jun 5;219(3):511-23.
12. Hahn CS, Strauss JH. Site-directed mutagenesis of the proposed catalytic amino acids of the Sindbis virus capsid protein autoprotease. Journal of Virology. 1990 Jun 1;64(6):3069-73.
13. Cen Y, Singh W, Arkin M, Moody TS, Huang M, Zhou J, Wu Q, Reetz MT. Artificial cysteine-lipases with high activity and altered catalytic mechanism created by laboratory evolution. Nature Communications. 2019 Jul 19;10(1):1-0.
14. Yan C, Du H. Lysosomal acid lipase is critical for myeloid-derived suppressive cell differentiation, development, and homeostasis. World Journal of Immunology. 2014 Jul 27;4(2):42-51.
15. Ogasawara D, Ichu TA, Vartabedian VF, Benthuysen J, Jing H, Reed A, Ulanovskaya OA, Hulce JJ, Roberts A, Brown S, Rosen H. Selective blockade of the lyso-PS lipase ABHD12 stimulates immune responses in vivo. Nature Chemical Biology. 2018 Dec;14(12):1099-108.
16. Flaherty SE, Grijalva A, Xu X, Ables E, Nomani A, Ferrante AW. A lipase-independent pathway of lipid release and immune modulation by adipocytes. Science. 2019 Mar 1;363(6430):989-93.
17. Cao J, Dang G, Li H, Li T, Yue Z, Li N, Liu Y, Liu S, Chen L. Identification and characterization of lipase activity and immunogenicity of LipL from Mycobacterium tuberculosis. PloS One. 2015;10(9).
18. Basketter D, Berg N, Kruszewski FH, Sarlo K, Concoby B. The toxicology and immunology of detergent enzymes. Journal of Immunotoxicology. 2012 Sep 1;9(3):320-6.
19. Girod A, Wobus CE, Zádori Z, Ried M, Leike K, Tijssen P, Kleinschmidt JA, Hallek M. The VP1 capsid protein of adeno-associated virus type 2 is carrying a phospholipase A2 domain required for virus infectivity. Journal of General Virology. 2002 May 1;83(5):973-8.
20. Dennis EA. The growing phospholipase A2 superfamily of signal transduction enzymes. Trends in biochemical sciences. 1997 Jan;22(1):1-2.
21. Afonso CL, Tulman ER, Lu Z, Oma E, Kutish GF, Rock DL. The genome of Melanoplus sanguinipes entomopoxvirus. Journal of Virology. 1999 Jan 1;73(1):533-52.
22. Deng H, Li W. Monoacylglycerol lipase inhibitors: modulators for lipid metabolism in cancer malignancy, neurological and metabolic disorders. Acta Pharmaceutica Sinica B. 2020 Apr 1;10(4):582-602.
23. Worley NB, Varela JA, Gaillardetz GP, Hill MN, Christianson JP. Monoacylglycerol lipase alpha inhibition alters prefrontal cortex excitability and blunts the consequences of traumatic stress in rat. Neuropharmacology. 2020 Jan 16:107964.
24. Thompson KJ, Tobin AB. Crosstalk between the M1 muscarinic acetylcholine receptor and the endocannabinoid system: A relevance for Alzheimer’s disease?. Cellular Signalling. 2020 Jan 21:109545.
25. Su H, Ruan YT, Li Y, Chen JG, Yin ZP, Zhang QF. In vitro and in vivo inhibitory activity of taxifolin on three digestive enzymes. International Journal of Biological Macromolecules. 2020 May 1;150:31-7.
26. Nasab SB, Homaei A, Pletschke BI, Salinas-Salazar C, Castillo-Zacarias C, Parra-Saldívar R. Marine resources effective in controlling and treating diabetes and its associated complications. Process Biochemistry. 2020 May 1;92:313-42.
27. Uppenberg J, Hansen MT, Patkar S, Jones TA. The sequence, crystal structure determination and refinement of two crystal forms of lipase B from Candida antarctica. Structure. 1994 Apr 1;2(4):293-308.
28. Stauch B, Fisher SJ, Cianci M. Open and closed states of Candida antarctica lipase B: protonation and the mechanism of interfacial activation. Journal of Lipid Research. 2015 Dec 1;56(12):2348-58.
29. Qu G, Li A, Sun Z, Acevedo-Rocha CG, Reetz MT. The crucial role of methodology development in directed evolution of selective enzymes. Angewandte Chemie International Edition. 2019 Jul 2.
30. Wang Y, Yu X, Zhao H. Biosystems design by directed evolution. AIChE Journal. 2020 Mar;66(3):e16716.
31. Arnold FH. Innovation by evolution: Bringing new chemistry to life (Nobel Lecture). Angewandte Chemie International Edition. 2019 Oct 7;58(41):14420-6.
32. Zeymer C, Hilvert D. Directed evolution of protein catalysts. Annual Review of Biochemistry. 2018 Jun 20;87:131-57.
33. Reetz MT. Laboratory evolution of stereoselective enzymes: a prolific source of catalysts for asymmetric reactions. Angewandte Chemie International Edition. 2011 Jan 3;50(1):138-74.
34. Acevedo-Rocha CG, Hoebenreich S, Reetz MT. Iterative saturation mutagenesis: a powerful approach to engineer proteins by systematically simulating Darwinian evolution. InDirected Evolution Library Creation 2014 (pp. 103-128). Springer, New York, NY.
35. Casa-Godoy, L, Gasteazoro, F, Bordes, F. Lipases: An Overview: Methods and Protocols. Methods in Molecular Biology 2018; 1835: 3-38.
36. Chahiniana H, Sarda L. Distinction between esterases and lipases: comparative biochemical properties of sequence-related carboxylesterases. Protein and peptide letters. 2009 Oct 1;16(10):1149-61.
37. Ali YB, Verger R, Abousalham A. Lipases or esterases: does it really matter? Toward a new bio-physico-chemical classification. InLipases and phospholipases 2012 (pp. 31- 51). Humana Press.
38. Bracco P, van Midden N, Arango E, Torrelo G, Ferrario V, Gardossi L, et al. Bacillus subtilis lipase a— lipase or esterase?. Catalysts. 2020 Mar;10(3):308.
