Abstract
Free-living amoebae are distributed worldwide and are found in a variety of environments. While most Acanthamoebae have been isolated from soil and water, A. royreba and A. culbertsoni were isolated from mammalian cell cultures. A. royreba’s isolation from malignant placental cell cultures, its ability to grow in mammalian cell culture media at 35oC, and containing a primitive centriole, prompted us to investigate the potential for mammalian information in A. royreba. While some soil amoeba contain small amounts of fungal and bacterial information, presumably from the microbes they phagocytosed, the informational content of A. royreba, in some instances, was very different with much of the mRNA being non-amoebic. Here we show that the proteins and mRNA content associated with A. royreba are extremely diverse and represent multiple Kingdoms, Orders, Phyla, and Genera from around the globe. The information in A. royreba, such as placental proteins from numerous mammals, the preponderance of non-amoebic mRNA, and its ability to tolerate harsh environments including Megarad irradiation leads to a discussion regarding the possible role of this amoebae as an immunological gatekeeper protecting fetal or malignant tissue from destruction and its potential as a vehicle for panspermia.
Introduction
Acanthamoeba was first described by Castellani [1] and represents single-cell eukaryotes existing as either cellular trophozoites (25–40 μm) or under adverse conditions (desiccation, lack of food, and extreme pH or temperature fluctuations) [2,3], as dormant cysts (13–20 μm). The cysts are known to be resistant to antibiotics, the effects of chlorine, and very low temperatures [4-6], and have been shown to maintain viability for over 20 years [7]. Acanthamoeba spp. have been placed into three groups (I, II, and III) based on the morphology and size of their cysts [8,9]. The 21 known Acanthamoeba spp. are ubiquitous and have been isolated from diverse environments including soil, water, compost, marine sediments, shellfish beds, and sludges [10-16]. They have also been isolated from fish, reptiles, and amphibians [17,18]. Two group III species, specifically A. culbertsoni and A. royreba, have been isolated from mammalian cell cultures [19,20]. Several have been implicated in a variety of diseases such as keratitis [21], encephalitis [22], and sinusitis [23].
Relative to a study we were undertaking regarding gallium-67 (67Ga) uptake in mammalian placentas and tumors [24,25], we were interested in pursuing the interaction of 67Ga with cultured human choriocarcinoma cells. Consequently, we obtained a culture of BeWo cells from Dr. Pattillo at the Medical College of Wisconsin (Milwaukee, Wisconsin, USA). This was the first human trophoblastic endocrine cell type to be maintained in continuous culture. We grew the culture in a mammalian cell culture medium (Plus I) with 10% added fetal calf sera (FCS). Shortly after obtaining this culture, events occurred which necessitated our laboratory being vacated for several weeks. In the interim, the BeWo cells had not received any media changes and subsequently experienced detrimental effects. While a majority of the BeWo cells were dead and floating upon our return, some of them were still attached and appeared viable. Surprisingly, some amoeboid motile cells were also observed intermixed with the few surviving.
BeWo cells. We sub-cultured the living amoeboid cells and sent them to Dr. Willaert at the Department of Protozoology, Institute of Tropical Medicine, Antwerp, Belgium. He observed that the locomotion of the cells was by protrusion of the cell cytoplasm involving the formation of pseudopodia with the creation and resorption of numerous spiked filiform projections. Under microscopic examination he determined that the amoebae were approximately 12–25 μm in length with a diameter of 10–15 μm. The amoebae also had a primitive centriole, relatively small nucleus with a single centrally located nucleolus, and abundant vacuoles. When the cells were placed in water they did not flagellate and when placed in saline or in media without added sera, the amoebae formed cysts typical of Acanthamoebae. Ultimately, he determined that the amoeboid cells were a new species: Acanthamoeba royreba [20,26].
While uncommon, ‘infection’ or ‘contamination’ of cultured mammalian cells (primary monkey kidneys cells, HeLa cells, and chick embryo cells) with free-living Acanthamoeba has been reported in the literature [19,27-29], with the first of these reports discussing the infection of cultured monkey kidney cells by an unidentified amoebae [30].
In light of the occasional unplanned sporadic isolation of free-living amoeba from cultured mammalian cells, including the unexpected isolation of A. royreba from BeWo cells, we deliberately attempted to induce the emergence of Acanthamoebae from various mammalian cell cultures by starvation. The untransformed cell lines tested included human W1-38 and mouse A-31 cell cultures. Malignant cell lines tested included rat hepatoma H35 and virus-transformed A31 Balb-C (KBALB) mouse cultures.
Plus I supplemented with 10% FCS was used as the base medium for the cell cultures which were starved (no medium exchanges) for 4–20 weeks. Visual and electron microscopic examination of the cultures after the starvation period did not reveal any evidence of amoebic cysts or trophozoites. While the addition of American Type Culture Collection (ATCC, Manassas, VA, USA) casitone medium #712 (typically used for Acanthamoeba cultures) to the flasks containing the starved cultures generally produced no growth, a few of the KBALB and H35 flasks did show the appearance of amoeboid cells after several attempts. The flasks that contained only the Plus I medium and FCS did not reveal the appearance of amoeboid cells nor did any of the A-31 or W1-38 cell cultures. Similar to the A. royreba culture isolated from the BeWo culture, the amoeboid cells from the KBALB and H35 cultures had characteristics (bulls-eye nucleolus, pseudopods, formed cysts, did not flagellate, and had vacuoles) typical of Acanthamoeba and were confirmed as such using immunological and serological assays. Dr. Willaert subsequently identified these isolates as A. culbertsoni[26].
All of the isolations of Acanthamoeba were difficult and required careful manipulation, observation, and the use of selective culture media. Ultimately, we felt that a more sophisticated approach to detect Acanthamoeba in placental or malignant tissue would be by the use of monoclonal antibodies. With some difficulty, a monoclonal antibody against A. royreba was produced [31].
Results and Discussion
A. culbertsoni was first isolated from monkey kidney cells [19], and the isolation of A. royreba occurred when human choriocarcinoma cells [32], were stressed by not changing the liquid culture media for several weeks. Similarly, when we deliberately starved separate cultures of malignant rat and mouse cells, we were also able to isolate cultures of A. culbertsoni [19]. The identification of A. royreba and A. culbertsoni from four separate axenic mammalian cultures prompted further investigations to understand the possible interaction between Acanthamoeba and other prokaryotes/eukaryotes. During our initial foray into these studies, we were curious whether phagocytic A. royreba would support replication of Legionella bacteria in manner similar to human macrophages; therefore, we deliberately infected A. royreba with an axenic culture of Legionella pneumophila [33], a Gram-negative bacterium that causes legionellosis [34]. While copious amounts of Legionella were replicated in A. royreba and subsequently cultured on Buffered Charcoal Yeast Extract medium, small amounts of diverse, somewhat unusual bacterial also grew on the Legionella-specific culture medium and tended to have unusual properties (highly pigmented and the ability to produce biogenic crystals) which differed from their conventional counterparts [35]. These bacteria were not culturable from the amoebae in the absence of the stress caused by the Legionella infection. This led to subsequent experiments using a ‘sterile stressor’, 1 Megarad gamma radiation, to further stress the amoeba with even more provocative results. As with the Legionella infection, previously unculturable bacteria could now be grown post-radiation [35]. In addition to biogenic crystal and pigment formation, many also produced extremely powerful biodispersants (capable of emulsifying mercury) and showed a diminished ability to ferment common sugars. Perhaps most interesting was that not only did the amoebic trophozoites survive the radiation dosage, but the bacteria that arose after gamma radiation were also radiation resistant! While stressing A. royreba resulting in the appearance of culturable bacteria mimics the stressing of the choriocarcinoma cells to yield culturable A. royreba in a Lego-like-linkage, the more interesting aspect of the radiation experiment was the ‘lethal/sterilizing’ Megarad dosage of gamma radiation used.
Having studied the interaction of A. royreba with Legionella which revealed the presence of additional prokaryotic bacteria, we then explored the possible sequestration of eukaryotic information in these amoebae since it exhibited some enigmatic mammalian characteristics. Not only was it isolated from human choriocarcinoma cells, but it could grow in several media used specifically for mammalian cell cultures and at temperatures as high as 35o C. Additionally, the cysts from this species could survive temperatures as low as -850 C. Unlike most other Acanthamoeba, A. royrebaand A. culbertsonihave primitive centrioles [20], an organelle which plays a major role in microtubule organization during cell division in other eukaryotes. Most importantly, monoclonal antibody production againstA. royreba, while easily accomplished in male mice, could not be accomplished using female mice. Interestingly, monoclonal antibodies that were produced against A. royreba interacted with a subpopulation of cells obtained from a monkey placental tissue (unpublished observations).
Initially, A. royreba was analyzed to detect the presence of mammalian proteins. Shotgun protein analysis was performed by Bioproximity (Manassas, VA, USA) [36-38]. The results indicated hundreds of mammalian and humanspecific proteins ranging from lymphocytes, erythrocytes, tumor rejection antigens, and sperm-binding proteins as well as several pregnancy-specific glycoproteins. While no one type commanded a high percentage of the total protein content, the diversity of these proteins was intriguing (Table 1). These results were interesting in that the protein information expressed in A. royreba contrasted with the genetic information in A. castellani as eloquently shown by Clarke et al. [39]. The genome of A. castellani showed the presence of predominantly Acanthamoeba genes and bacterial and fungal information which amounted to approximately 2.9% of the genome, presumably from ingestion of these microbes, and could represent lateral gene transfer (LGT). LGT, or horizontal gene transfer [40,41], is the transmission of genes between individual cells (from species to species) and can generate new gene sequences through transformation, transduction, and conjugation.
| Archaea (0%) | Bacteria (37.69%) | Eukarya (62.31%) |
|---|---|---|
| 15 Phyla1 | Fungi – 3 Phyla2 | |
| Animalia – 4 Orders3 | ||
| Protista4 | ||
| 1 representative habitats include: marine, freshwater, brackish, sludge, sand, gut flora, soil 2 includes lichens 3 representative mammals include: humans, placental mammals, shrews, bats, even-toed hoofed animals, new world monkeys, African ruminant animals, cattle 4 includes 4 spp. of Acanthamoeba (A. castellanii: 99.99%) and Proteobacteria |
||
Subsequently, A. royreba, axenically cultured in a proteose peptone-yeast extract-glucose (PYG) medium with 40 μg gentamicin/ml [42], was tested to determine its messenger RNA (mRNA) content by both Quick Biology (Pasadena, CA, USA) and My Genomics (Atlanta, GA, USA).
The diversity shown in the protein analysis of A. royreba was further confirmed and greatly magnified in the content of the mRNAs (Tables 2 and 3, and Figures 1 and 2). Duplicate frozen cultures of A. royreba were thawed and grown at 37o C. The results showed that after 2 weeks approximately 25% of the mRNA profile was non-amoebic (Figure 1), which increased to approximately 70% after 2 months (Figure 2), indicating a possible time- and/or temperature-dependent genetic expression component. While not the norm, growing A. royreba at 37o C may be the road best traveled for revealing pivotal mRNA profiles. While less than half of the mRNA in the cells cultured for 2 months represented characteristics specific to Acanthamoeba, the remaining spectrum of mRNA content was very diverse and represented fungal, chordata, arthropoda, and bacterial communities. As noted in the protein analysis, approximately 3% of the mRNA was mammalian (Tables 2 and 3, and Figure 2).
| Eukarya 95.25% | Bacteria 4.46% | ||||
|---|---|---|---|---|---|
| Protista 37.85% |
Fungi 25.77% |
Animalia 22.41% | Plantae 4.75% | Other Eukaryotes 4.47% |
|
| Acanthamoebae 30.76% (A. castellani 26.81%) |
Ascomycota 14.75% |
Arthropodia 12.41% |
Proteobacteria 0.28% |
||
| Algae 7.09% |
Basidiomycota 11.02% |
Chordata 10.01% | Viruses 0.01% |
||
| Algae | 99 different Genera | Red, green, flagellated |
| Fungi | 360 different Genera | Mushrooms, yeasts, molds |
| Bacteria | 67 different Genera | Pathogens, nonpathogens, soil, water, hot springs |
| Insects | 120 different Genera | Mosquitos, flies, moths, butterflies, ants, wasps, bees, beetles, planthopper, bed bugs, mites, aphids, ticks |
| Fish | 41 different Genera | Coelacanth, whitefish, sablefish, minnow, lancelets, herrings, pupfish, carp, danios, pike, cichlid, catfish, croakers, gars, icefish, mollies, damselfish |
| Marine organisms and crustaceans |
26 different Genera | Crabs, lobsters, squid, sperm whale, sea squirt, prawn, sea slug, oysters, barnacles, tube worms, sponges |
| Mammals and amphibians |
52 different Genera | Mouse, camel, elephant, goat, alligator, lizard, Seriema, guinea pig, boar, turtles, crocodiles, bats, gecko, gorilla, squirrel, Weddell seal, turkey, hamster, lemurs, ferret, mole-rat, frogs, degu, aardvark, rabbit, baboons, shrews |
| Crops | 22 different Genera | Rice, corn, cucumber, soybean, apples, pears, radishes, castor bean, sesame |
| Birds | 11 different Genera | Pigeons, falcon, woodpeckers, starlings |
| Trees | 21 different Genera | Gum tree, locust tree, walnut tree, date palm, poplar, tropical evergreens |
| Plants | 17 different Genera | Rose, lupine, various grasses |
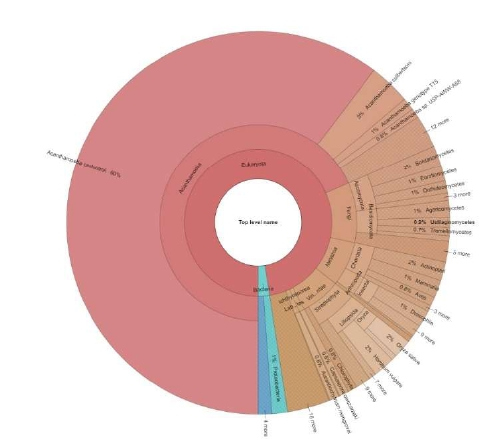
Figure 1: mRNA analysis of A. royreba grown for 2 weeks prior to analysis showing diversity of taxa
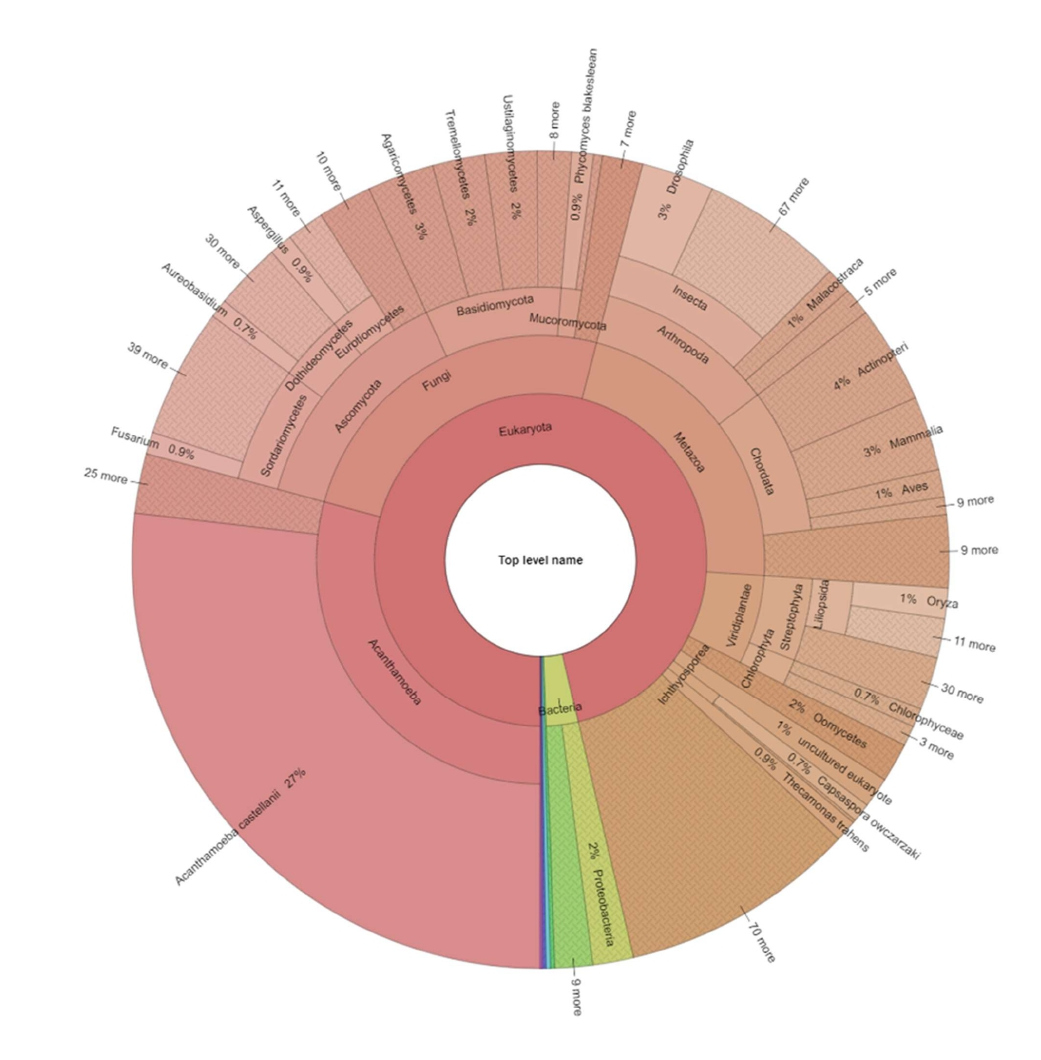
Figure 2: mRNA analysis of A. royreba grown for 2 months prior to analysis showing extreme diversity of taxa.
Since many Acanthamoebae live primarily in soil and water, ingest a variety of fungal and bacterial microbes, and contain bacterial endosymbionts, it is understandable that the amoebae contain mRNAs indicative of these organisms. However, the fact that there is a species, A. royreba, which contains mRNA specific to a wide and diverse spectra within multiple Kingdoms, Phyla, and Genera is both puzzling and fascinating and may have profound evolutionary implications.
The questions these experiments raise have profound implications in evolutionary biology, physiology, environmental science, medicine, exobiology, and paleobiology, to mention just a few. Several admittedly speculative implications arise from A. royreba being an immense reservoir of global genetic information:
(1) Is there some analogy to be drawn from the relationship between mammalian cells and an amoebae and the bacterial origin of mitochondria [43]?
(2) A. royreba was originally isolated from a malignant human placental trophoblast (i.e., choriocarcinoma), contains placental proteins from various mammals, and was not recognized as foreign by female mice, but was recognized as such by male mice. Also, the antibodies made against A. royreba did react with some cells from a Rhesus monkey placenta. Therefore, could these observations indicate that these amoebae exist as endosymbionts in placental or tumoral tissue? Could these amobae possibly play a crucial role in the womb becoming an immunologic sanctuary that permits the presence of a foreign fetus without rejection? If so, can this unusual property be extrapolated to explain the tolerance of the immune system to fetal or tumoral tissue?
Interestingly, when A. royreba was placed in a coculture of human red and white blood cells, the amoebae preferentially destroyed the white cells [19]. Similarly, the study by Chi et al. [28], also showed that the Acanthamoeba they isolated from monkey kidney cells would specifically attack and enucleate chick erythrocytes but would ignore guinea pig erythrocytes. These observations indicate a mechanism whereby amoebae can ‘identify’ different cell types. Obviously, a placenta-based amoebic ‘immunologic gate keeper’ would want to allow entry of red blood cells, but not potentially harmful white blood cells.
(3) A. royreba can tolerate temperatures of -85o C, MegaRad doses of gamma radiation, form cysts that contain and safeguard a vast spectrum of genetic information, and harbor or spawn numerous bacteria or bacterialike organisms with unusual properties in an apparent symbiotic-like relationship allowing both to survive harsh environments. Can it therefore be a candidate as a vehicle for panspermia thereby shedding light on the transfer of life [44,45]? Is it also imaginable that an entity such as A. royreba could, perhaps encased in asteroids, comets, or space dust, traverse many light years and seed planets in the cosmos – including Earth?
Conclusion
In conclusion, the results of this preliminary study show that A. royreba is an unrealized reservoir of complex genomic information that could be a primary candidate for understanding some of the greatest mysteries in biology. The ability of this unique organism to adapt to its environment and modify its surroundings makes A. royreba an organism worthy of further study. Now we look through the glass darkly, but the images are still very exciting.
Materials and Methods
Protein and mRNA analyses (accession number QD512)
Shotgun protein analysis was performed by Bioproximity: 9385 Discovery Blvd, Manassas, VA 20109; info@ bioproximity.com
mRNA analyses was performed by Quick Biology: 2265 E Foothill Blvd., Pasadena, CA, 91107 and My Genomics; Atlanta, GA, USA; info@mygenomics.com
RNA integrity was checked using an Agilent Bioanalyzer 2100 (Agilent, Santa Clara, CA, USA); only samples with clean rRNA peaks were used for further experiments. The RNA-Seq Library was prepared according to KAPA Stranded mRNA-Seq poly(A) selected kits with 201-300 bp insert sizes (KAPA Biosystems, Wilmington, MA, USA) using 250 ng total RNA as an input. Final library quality and quantity were analyzed using an Agilent Bioanalyzer 2100 and a Life Technologies Qubit 3.0 Fluorometer (ThermoFisher Scientific, Waltham, MA, USA). 150 bp paired-end reads were sequenced on an Illumina HiSeq 4000 (Illumnia Inc., San Diego, CA). 10,000 random reads were used for the blast searching against the NCBI nr database. A custom perl script was used for the taxonomy analysis based on the blast result. Krona was used to generate multi-layered pie charts.
Methods regarding the growth of discussed organisms
Amoebae were grown following the procedure of Willaert et al. [20].
Choriocarcinoma cells were cultured as described in Pattile et al. [32].
Monoclonal antibody production in mice used the procedure of Kennel et al. [31].
Gamma irradiation procedures and the growth and isolation of endosymbionts followed the protocol outlined in Vass et al. [35].
Legionella were cultured as described in Tyndall et al. [33] and Lau et al. [34].
A. royreba, was axenically cultured in a proteose peptone-yeast extract-glucose (PYG) medium with 40 μg gentamicin/ml [42].
Acknowledgments
We thank Audrey Nicholson who first detected A.royreba in the choriocarcinoma cell culture and Shantanu Roy for assistance in sample preparation. Quick Biology, Pasadena, CA, provided Figures 1 and 2.
Disclaimer
The findings and conclusions in this report are those of the authors and do not necessarily represent the official position of the Centers for Disease Control and Prevention.
References
2. Byers TJ, Akins RA, Maynard BJ, Lefken RA, Martin SM. Rapid growth of Acanthamoeba in defined media; induction of encystment by glucose-acetate starvation. The Journal of protozoology. 1980 May;27(2):216-9.
3. Chagla AH, Griffiths AJ. Growth and encystation of Acanthamoeba castellanii. Microbiology. 1974 Nov 1;85(1):139-45.
4. Khunkitti W, Lloyd DF, Furr JR, Russell AD. Acanthamoeba castellanii: growth, encystment, excystment and biocide susceptibility. Journal of Infection. 1998 Jan 1;36(1):43-8.
5. De Jonckheere J, Van de Voorde H. Differences in destruction of cysts of pathogenic and nonpathogenic Naegleria and Acanthamoeba by chlorine. Applied and Environmental Microbiology. 1976 Feb 1;31(2):294-7.
6. Lloyd D, Turner NA, Khunkitti W, Hann AC, Furr JR, Russell AD. Encystation in Acanthamoeba castellanii: Development of Biocide Resistance 1. Journal of Eukaryotic Microbiology. 2001 Jan;48(1):11-6.
7. Mazur T, Hadas E, Iwanicka I. The duration of the cyst stage and the viability and virulence of Acanthamoeba isolates. Tropical medicine and parasitology: official organ of Deutsche Tropenmedizinische Gesellschaft and of Deutsche Gesellschaft fur Technische Zusammenarbeit (GTZ). 1995 Jun;46(2):106-8.
8. Visvesvara GS. Classification of Acanthamoeba. Reviews of Infectious Diseases. 1991 Mar 1;13(Supplement_5):S369-72.
9. Pussard M. Morphologie de la paroi kystique et taxonomie du genre Acanthamoeba (Protozoa, Amoebida).
10. Todd CD, Reyes-Batlle M, Martín-Navarro CM, Dorta-Gorrín A, López-Arencibia A, et al. Isolation and genotyping of Acanthamoeba strains from soil sources from Jamaica, West Indies. Journal of Eukaryotic Microbiology. 2015 May;62(3):416-21.
11. Sawyer TK, Visvesvara GS, Harke BA. Pathogenic amoebas from brackish and ocean sediments, with a description of Acanthamoeba hatchetti, n. sp. Science. 1977 Jun 17;196(4296):1324-5.
12. Page FC. Re-definition of the genus Acanthamoeba with descriptions of three species. The Journal of Protozoology. 1967 Nov;14(4):709-24.
13. Xuan Y, Shen Y, Ge Y, Yan G, Zheng S. Isolation and identification of Acanthamoeba strains from soil and tap water in Yanji, China. Environmental Health and Preventive Medicine. 2017 Dec 1;22(1):58.
14. Lewis EJ, Sawyer TK. Acanthamoeba tubiashi N. Sp., a new species of fresh-water Amoebida (Acanthamoebidae). Transactions of the American Microscopical Society. 1979 Oct 1:543-9.
15. Sawyer TK, Buchanan LR. Contamination of tissue sections of the American oyster by cysts of Acanthamoeba sp. Journal of Invertebrate Pathology. 1971 Sep;18(2):300.
16. Sawyer TK, Nerad TA, Lewis EJ, Mclaughlin SM. Acanthamoeba stevensoni N. Sp.(Protozoa: Amoebida) from Sewage-Contaminated Shellfish Beds in Raritan Bay, New York. Journal of Eukaryotic Microbiology. 1993 Nov;40(6):742-6.
17. Sesma MJ, Ramos LZ. Isolation of free-living amoebas from the intestinal contents of reptiles. The Journal of Parasitology. 1989 Apr 1:322-4.
18. Dykova I, Lom J, Schroeder-Diedrich JM, Booton GC, Byers TJ. Acanthamoeba strains isolated from organs of freshwater fishes. The Journal of Parasitology. 1999 Dec 1:1106-13.
19. Culbertson CG, Smith JW, Minner JR. Acanthamoeba: observations on animal pathogenicity. Science. 1958 Jun 27;127(3313):1506.
20. Willaert E, Stevens AR, Tyndall RL. Taxonomy Acanthamoeba royreba sp. n. from a Human Tumor Cell Culture. The Journal of Protozoology. 1978 Feb;25(1):1-14.
21. Seal D, Hay J, Kirkness C, Morrell A, Booth A, Tullo A, et al. Successful medical therapy of Acanthamoeba keratitis with topical chlorhexidine and propamidine. Eye. 1996 Jul;10(4):413-21.
22. Marciano-Cabral FR, Bradley SG. Current research on free-living amebae causing granulomatous amebic encephalitis. The Journal of Eukaryotic Microbiology. 2001 Jan;48(7):3-4.
23. Rivera MA, Padhya TA. Acanthamoeba: a rare primary cause of rhinosinusitis. The Laryngoscope. 2002 Jul;112(7):1201-3.
24. Swartzendruber DC, Byrd BL, Hayes RL, Nelson B, Tyndall RL. Preferential localization of gallium-67 citrate in tissues of leukemic mice. Journal of the National Cancer Institute. 1970 Mar 1;44(3):695-700.
25. Tyndall RL, Chaskes SJ, Carlton JE, Nelson B, Daniel JC. Gallium-67 distribution in pregnant mammals. Journal of Experimental Zoology. 1976 Mar;195(3):417- 23.
26. Tyndall RL. Pathogenic and enzymatic characteristics of Acanthamoeba isolated from cultured tumor cells. Protistologica. 1979;1:17-22.
27. Casemore DP. Contamination of virological tissue cultures with a species of free-living soil amoeba. Journal of Clinical Pathology. 1969 May 1;22(3):254-7.
28. Chi L, Vogel JE, Shelokov A. Selective phagocytosis of nucleated erythrocytes by cytotoxic amebae in cell culture. Science. 1959 Dec 25;130(3391):1763-4.
29. Dux K. An intracellular agent inducing hypertrophy and hyperplasis in chick embryo cultures. Archivum Immunologiae et Therapiae Experimentalis. 1969;17:624- 32.
30. Jahnes WG, Fullmer HM, Li CP. Free living amoebae as contaminants in monkey kidney tissue culture. Proceedings of the Society for Experimental Biology and Medicine. 1957 Nov;96(2):484-8.
31. Kennel SJ, Foote LJ, Lankford PK. Analysis of surface proteins of mouse lung carcinomas using monoclonal antibodies. Cancer Research. 1981 Sep 1;41(9 Part 1):3465- 70.
32. Pattillo RA, Gey GO. The establishment of a cell line of human hormone-synthesizing trophoblastic cells in vitro. Cancer Research. 1968 Jul 1;28(7):1231-6.
33. Tyndall RL, Domingue EL. Cocultivation of Legionella pneumophila and free-living amoebae. Applied and Environmental Microbiology. 1982 Oct 1;44(4):954-9.
34. Lau HY, Ashbolt NJ. The role of biofilms and protozoa in Legionella pathogenesis: implications for drinking water. Journal of Applied Microbiology. 2009 Aug;107(2):368-78.
35. Vass AA, Mackowski RP, Tyndall RL. Appearance of viable bacteria in Acanthamoeba royreba after amoebic exposure to megarad doses of gamma radiation. InEukaryotism and Symbiosis 1997 (pp. 379-388). Springer, Berlin, Heidelberg.
36. Rappsilber J, Mann M, Ishihama Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nature Protocols. 2007 Aug;2(8):1896-1906.
37. Chambers MC, Maclean B, Burke R, Amodei D, Ruderman DL, Neumann S, Gatto L, Fischer B, Pratt B, Egertson J, Hoff K. A cross-platform toolkit for mass spectrometry and proteomics. Nature Biotechnology. 2012 Oct;30(10):918-20.
38. Wisniewski JR, Zougman A, Nagaraj N, Mann M. Universal sample preparation method for proteome analysis. Nature Methods. 2009 May;6(5):359-62.
39. Clarke M, Lohan AJ, Liu B, Lagkouvardos I, Roy S, Zafar N, Bertelli C, Schilde C, Kianianmomeni A, Bürglin TR, Frech C. Genome of Acanthamoeba castellanii highlights extensive lateral gene transfer and early evolution of tyrosine kinase signaling. Genome Biology. 2013 Feb;14(2):1-5.
40. Dunning Hotopp JC. Horizontal gene transfer between bacteria and animals. Trends in Genetics. 2011 Apr 1;27(4):157-63.
41. Ochman H, Lawrence JG, Groisman EA. Lateral gene transfer and the nature of bacterial innovation. Nature. 2000 May;405(6784):299-304.
42. Visvesvara GS, Balamuth W. Comparative studies on related free-living and pathogenic amebae with special reference to Acanthamoeba. The Journal of Protozoology. 1975 May;22(2):245-56.
43. Margulis L, Sagan D. Origins of Sex. Three Billion Years of Genetic Recombination. Yale University Press: New Haven; 1986. pp. 69–71.
44. Crick FH, Orgel LE. Directed panspermia. Icarus. 1973 Jul 1;19(3):341-8.
45. Wickramasinghe C. Bacterial morphologies supporting cometary panspermia: a reappraisal. International Journal of Astrobiology. 2011;10(1):25-30.
